Giornata di presentazione della rete di Biologia Strutturale del CNR (SBN@CNR), Roma, 16 Aprile 2025
Consiglio Nazionale delle Ricerche - Aula Marconi
Piazzale Aldo Moro 7 - Roma
Cari colleghi e care colleghe,
Il convegno da noi organizzato ha lo scopo di creare un network di biologia strutturale (SBN-CNR) per potenziare l’impatto della biologia strutturale del CNR, perché possa essere di aiuto a tutta la comunità scientifica italiana nella risoluzione di problemi biologici e nella progettazione di nuovi “lead compound” attraverso lo “Structure Based Drug Design”. La diffusione di tecniche innovative come la “Cryo-EM” e la possibilità di predire le strutture di proteine sfruttando programmi di intelligenza artificiale come “AlphaFold”, hanno rivoluzionato la biologia strutturale. L'espansione delle conoscenze strutturali richiede nuovi approcci e una cooperazione interdisciplinare, missione centrale della rete SBN@CNR.
Speriamo di incontrarvi numerosi alla giornata.
Il comitato organizzatore
Link evento: https://sites.google.com/view/rete-sbn-cnr
Link programma: https://sites.google.com/view/rete-sbn-cnr/programma
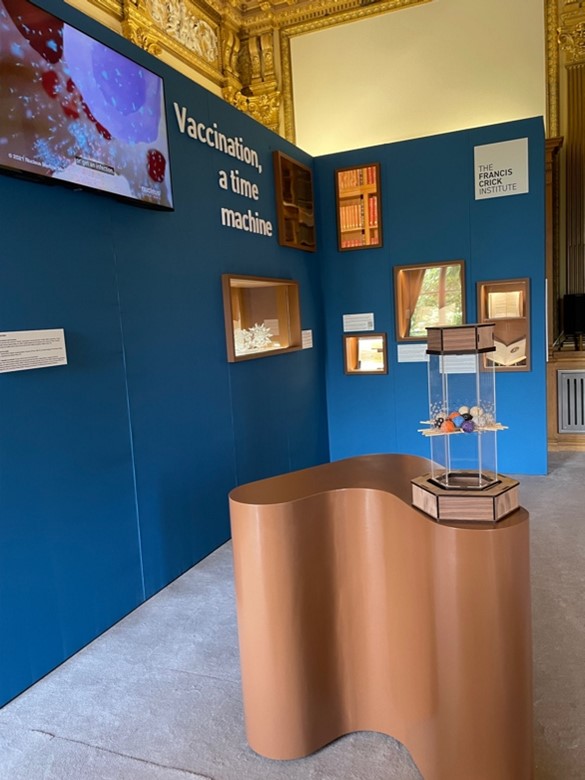
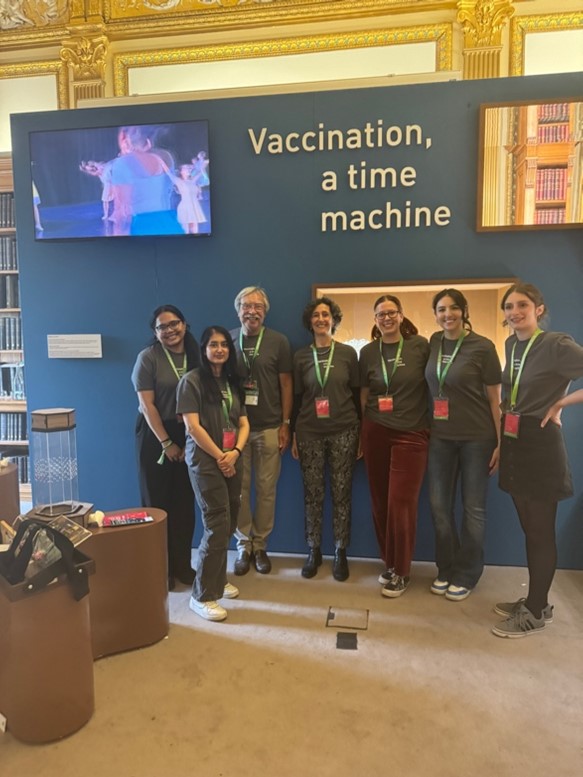
"Vaccination, a time machine", exhibition, July 2024

is the title of the exhibition that Francesca Di Rosa, IBPM Research Director and Visiting Scientist at the Francis Crick Institute in London, curated for the 2024 edition of the prestigious Royal Society Summer Exhibition, in collaboration with Prof Adrian Hayday (Francis Crick Institute and King's College London).
To engage visitors, Francesca chose a multidisciplinary approach, as she herself explains in the article just published in Nature Review Immunology entitled “Using art and history to communicate immunology to a broad audience”, accessible here https://rdcu.be/dT5KH


3/10/2024
IBPM-CNR @ the European Biotech Week 2024 (EBW2024)
"DNA HUNT: INVESTIGATING THE CRIME SCENE"
Giovedì 3 Ottobre , 10 :00-14:00 - Sede Centrale CNR - Aula Marconi, piazzale Aldo Moro 7, Roma
A detective day for secondary school classes that offers a simulated experience of 'DNA profiling' used in forensic laboratories. A virtual tour will guide students through the key stages of understanding the genome.

Organizzazione evento IBPM-CNR per la EBW2024: Teresa Colombo, Giovanna Costanzo, Alessandra Guidi
Link: https://www.cnr.it/it/evento/19381
Link: https://assobiotec.federchimica.it/agenda/tutti-gli-eventi/2024/09/24/default-calendar/biotech-week-2024
27/09/2024
IBPM-CNR @ the European Researchers' Night 2024 (ERN2024)
EVENT:
"DNA take away"
27 September - 28 September 2024, 18:30-23:00 - Città dell'Altra Economia, Testaccio, Roma
Largo Dino Frisullo, snc, 00153 Roma RM
Free entry event, for ages 8+. Note: Online reservation is required on the website https://www.scienzainsieme.it/ (The direct link for the DNA Take Away event is: https://www.scienzainsieme.it/dna_take-away/) to participate in the experimental activity.
A scientist's experiment! Participants will be involved in a special experiment, simple but scientifically rigorous, which will allow them to extract their DNA and create a "bow" and then put in a bottle. What to do next? Take it with you, to remind yourself how amazing science is!

Link: https://www.scienzainsieme.it/notte-europea-dei-ricercatori/
22/11/2023
Nel DNA della popolazione nomade dei Fulani la testimonianza di antichi pastori sahariani
Uno studio, condotto da un team internazionale coordinato da Sapienza Università di Roma e dall’Istituto di biologia e patologia molecolari del Consiglio nazionale delle ricerche, mostra un’affinità genetica tra Fulani e individui del Marocco risalenti al Neolitico. I risultati, pubblicati sulla rivista Current Biology, individuano nei pastori nomadi Fulani i discendenti di una antica po-polazione “pan-Sahariana” che si è poi definitivamente frammentata e divisa con la desertificazio-ne della regione africana
Eugenia D'Atanasio eugenia.datanasio@cnr.it
Fulvio Cruciani fulvio.cruciani@uniroma1.it
Un team di ricerca internazionale, coordinato dalla Sapienza e dall’Istituto di biologia e patologia molecolari del Consiglio nazionale delle ricerche (Cnr-Ibpm) di Roma, ha condotto uno studio che fa luce sul ruolo dei cambiamenti climatici del Sahara.
Sfruttando l’informazione contenuta nell’intero genoma di 43 individui e applicando le nuove tecnologie di sequenziamento ad alta copertura, i ricercatori si sono focalizzati sui Fulani, la più estesa popolazione di pastori nomadi al mondo, attualmente stanziata lungo tutta la fascia saheliana, immediatamente a sud del deserto del Sahara. Questa popolazione, da studi genetici e antropologici precedenti, sembrava presentare delle caratteristiche peculiari, diverse da quelle delle popolazioni circostanti.
Lo studio, pubblicato sulla rivista Current Biology, ha indagato la natura di questa specificità al fine di darne anche una caratterizzazione storica e cronologica attraverso il confronto con altri genomi moderni e antichi presenti in letteratura.
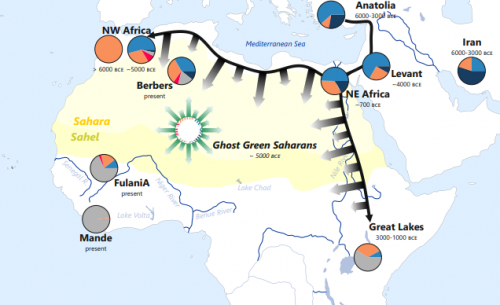
Rappresentazione semplificata delle componenti genetiche nelle popolazioni del Green Sahara e nei Fulani.
(Credits: Eugenia D'Atanasio. Concessione di Current Biology)
Per la prima volta è stata evidenziata una forte affinità genetica, finora solo ipotizzata sulla base di elementi culturali e reperti archeologici, tra i Fulani e individui del Marocco risalenti al Neolitico. Con un approccio di analisi congiunta di genomi moderni e antichi mai applicato per il continente africano, è stato possibile collocare l’origine dei Fulani nell’arco dell’ultimo periodo umido del Sahara (12.000 – 5.000 anni fa, denominato “Green Sahara”), quando l’attuale deserto era invece una terra fertile, rigogliosa e abitata, facendo di questa popolazione la testimone chiave di uno dei più grandi cambiamenti climatici del continente africano.
“Gli approcci utilizzati in questo studio ci hanno inoltre permesso di fare luce sulla storia recente dei Fulani, che risultano suddivisi in due sotto-gruppi geneticamente distinti a causa di fattori socio-culturali non ancora ben caratterizzati”, spiega Flavia Risi, dottoranda in genetica e biologia molecolare presso la Sapienza Università di Roma e co-autrice della ricerca.
Lo scenario ipotizzato sulla base di questi risultati propone inoltre la presenza nel “Green Sahara” di frequenti contatti e migrazioni tra le popolazioni, a formare una sorta di popolazione “pan-Sahariana” che si è poi definitivamente frammentata e divisa con la desertificazione dell’area, a partire da 5.000 anni fa ma di cui rimane traccia nei Fulani in virtù di uno stile di vita particolarmente isolato.
“L’ambiente fertile del Sahara ha sicuramente promosso estesi movimenti e contatti tra le popolazioni del Sahara e i risultati di questo studio aiutano a tracciare una linea di congiunzione tra gli antichi pastori sahariani e i moderni Fulani” spiega Eugenia D’Atanasio, ricercatrice dell’Istituto di biologia e patologia molecolari del Cnr (Cnr-Ibpm) e co-coordinatrice dello studio.
Lo studio rappresenta un contributo alle conoscenze attuali sulla storia evolutiva umana e sulla sua biodiversità, in particolare nel continente africano, luogo che ospita la più grande ricchezza genetica per la nostra specie. Tali risultati evidenziano poi come spostamenti e contatti transahariani non siano solo un fenomeno recente, ma una realtà storica antichissima, confermando infine il ruolo dei grandi cambiamenti climatici (come la desertificazione) nelle migrazioni umane.
“L’Africa, nonostante la sua ricchezza culturale e genetica, è ancora poco studiata a livello genomico. Oltre alle informazioni sul popolamento del “Green Sahara”, le sequenze genomiche prodotte durante questo studio rappresentano uno strumento essenziale per la comunità scientifica in quanto permettono di colmare, almeno in parte, una grossa lacuna nella nostra conoscenza della diversità umana”, aggiunge Fulvio Cruciani del Dipartimento di Biologia e biotecnologie Charles Darwin della Sapienza Università di Roma e PI del progetto.
18/09/2022
IBPM-CNR @ the European Researchers' Night 2023 (ERN2023)
EVENT:
"DNA take away"
30 September - 01 October 2022, 18:30-23:00 - Città dell'Altra Economia, Testaccio, Roma
Evento ad accesso gratuito, ma è consigliata la PRENOTAZIONE
A scientist's experiment! Participants will be involved in a special experiment, simple but scientifically rigorous, which will allow them to extract their DNA and create a "bow" and then put in a bottle. What to do next? Take it with you, to remind yourself how amazing science is!

Link: https://www.scienzainsieme.it/notte-europea-dei-ricercatori/
18/09/2022
IBPM-CNR @ the European Biotech Week 2023 (EBW2023)
EVENT 1:
"A walk into your genome"
28 September, 9:00-14:00 - Sede Centrale CNR - Aula Marconi, piazzale Aldo Moro 7, Roma
A day of exploration of our genome dedicated to upper secondary school students. We will recount the milestones in our knowledge of the human genome, and how this achievement has changed the face of bio-medical research, our view of the organization of our genome, and our understanding of the mechanisms underlying many diseases. We will take a virtual tour of the human genome using a kind of genomic Google Maps. There will be quizzes designed to delve into fascinating numbers and facts about our genome. In addition to the presentations and tour stops, participants will have the exciting opportunity to extract their own DNA. What's even more exciting is that, for the first time in this edition, participants will have the valuable assistance of an extraordinary lab companion—an 'edudroid,' an interactive robot that's eager to answer questions and spark curiosity about DNA.
EVENT 2:
"A dip in the magic world of water!"
26 September 2023 (9:00-13:00); 27 September 2023 (9-16) - Scuola primaria dell'Istituto Comprensivo "Martin Luther King", via degli Orafi 30, Roma
Water has always represented a fascinating world of great fun for children. This workshop aims to explore the extraordinary properties of water, which we experience every day, and that will be explained to children in a playful but at the same time scientific way. The laboratory plans to explore together some common properties of water (temperature, state, capillarity, surface tension and density) using simple language and experiments aimed at understanding such characteristics.

Organizzazione evento IBPM-CNR per la EBW2022: Giovanna Costanzo, Angela Cirigliano, Teresa Colombo, Alessandra Guidi, Cecilia Mannironi, Erica Salvati
Link: https://www.cnr.it/it/evento/18667/il-cnr-ibpm-per-la-biotech-week-2023
Link: https://assobiotec.federchimica.it/agenda/tutti-gli-eventi/2023/09/25/default-calendar/biotech-week-2023
16/02/2023
Sharing the future - GIOVANI e SCIENZA

Organizzazione evento:
CNR Unità comunicazione
Fondazione Mondo DIgitale
Link CNR's page: https://www.cnr.it/it/evento/18381/sharing-the-future-giovani-e-scienza
Link to the program: Agenda_Sharing_the_future_CNR_sede_16febb23.pdf
19/09/2022
IBPM-CNR @ the European Biotech Week 2022 (EBW2022)
EVENT 1:
"A walk into your genome"
Monday 26 September, 9:00-13:00 - Sede Centrale CNR - Aula Marconi, piazzale Aldo Moro 7, Roma
A day of exploration of our genome meant with the students of the Liceo Ginnasio "T. Tasso" in Rome. We will recall the core steps towards knowledge of the human genome, and discuss how this achievement has changed the face of bio-medical research, the vision we have of the organization of our genome and the understanding of the mechanisms underlying many diseases. We will take a virtual tour of the human genome by using a sort of genomic Google Maps. There will be quizzes to explore numbers and surprising facts about our genome. Between a presentation and a stop on our tour, participants will be able to take part in a real extraction of their own DNA.
EVENT 2:
"A dip in the magic world of water!"
27-28-29-30 September 2022, 9:00-12:00 - Scuola primaria ed infanzia "Vittorio Piccinini", Istituto Comprensivo "Anna Fraentzel Celli", via F. Fiorentini 48, Roma
Water has always represented a fascinating world of great fun for children. This workshop aims to explore the extraordinary properties of water, which we experience every day, and that will be explained to children in a playful but at the same time scientific way. The laboratory plans to explore together some common properties of water (temperature, state, capillarity, surface tension and density) using simple language and experiments aimed at understanding such characteristics. The activity is offered to children of two different age groups (3-6 and 6-10) for which we will keep the theme while varying the modality and language to the age range of the participants.

Organizzazione evento IBPM-CNR per la EBW2022: Giovanna Costanzo, Angela Cirigliano, Teresa Colombo, Alessandra Guidi, Cecilia Mannironi, Erica Salvati
Link: https://www.cnr.it/it/evento/18077/il-cnr-ibpm-per-la-biotech-week-2022
Link: https://assobiotec.federchimica.it/agenda/tutti-gli-eventi/2022/09/26/default-calendar/biotech-week-2022
19/09/2022
IBPM-CNR @ the European Researchers' Night 2022 (ERN2022)
EVENTO:
"My DNA? Today I'll put it in a bottle"
30 September - 01 Ottobre 2022, 18:30-23:00 - Città dell'Altra Economia, Testaccio, Roma
Evento ad accesso gratuito, ma è consigliata la PRENOTAZIONE
A scientist's experiment! Participants will be involved in a special experiment, simple but scientifically rigorous, which will allow them to extract their DNA and create a "bow" and then put in a bottle. What to do next? Take it with you, to remind yourself how amazing science is!

Link: https://www.scienzainsieme.it/il-mio-dna-oggi-lo-metto-in-bottiglia/
Link: https://www.scienzainsieme.it/category/notte-2022/roma/
20/06/2022
The first complete sequenze of a human genome: IBPM talks about it on show L'Italia con Voi
On April 1, 2022, the journal Science published the work by the Telomere-2-Telomere (T2T) consortium that illustrates achievement of a new fundamental milestone in biomedicine: the first complete sequence - from telomere to telomere - of a human genome. Two IBPM researchers, Teresa Colombo and Giovanna Costanzo, comment on it in the TV talk "L'Italia con Voi" broadcasted by the RAI Italia channel on the last June the 14th, and now visible on the RaiPlay website at this link: https://www.raiplay.it/video/2022/06/LItalia-con-Voi-Puntata-del-14062022-16384347-0525-4f70-a9dc-20c25f2e9035.html (IBPM intervention at around minute 45')
01/02/2022
Treatment of kidney clear cell carcinoma, lung adenocarcinoma and glioblastoma cell lines with hydrogels made of DNA nanostars
Biomaterials Science
Link: https://pubs.rsc.org/en/content/articlelanding/2022/bm/d1bm01643a
Can DNA-GELs made of DNA nanostars help with the treatment of tumor cell lines?
17/09/2021
POR FESR 2014-2020 Avviso Pubblico “Gruppi di Ricerca - 2020” Progetti finanziati

APTASER - Sviluppo di inibitori ad acido nucleico della serina idrossimetiltrasferasi nel tumore al polmone
DWARF - La felce Pteris vittata: una strategia green per eliminare l’arsenico dalle acque potabili
INNOVA3DIMAGING - Metodologie innovative di 3D high content imaging per la validazione di nuove molecole antitumorali
NEWPAT - Analisi genetica del DNA fetale circolante nel sangue materno: sviluppo di un test di paternità non invasivo ad elevata specificità
ORGANOVA - Sviluppo di modelli organotipici e organ-on-chip di tumore ovarico per testare farmaci antitumorali
TOC - interazioni tra luce e ormoni nel controllo dell’apertura fiorale in specie modello e ortive
SMART-BREED - Tecnologie molecolari innovative per l’adattamento delle specie ortive al cambiamento climatico mediante breeding di precisione
27/07/2021
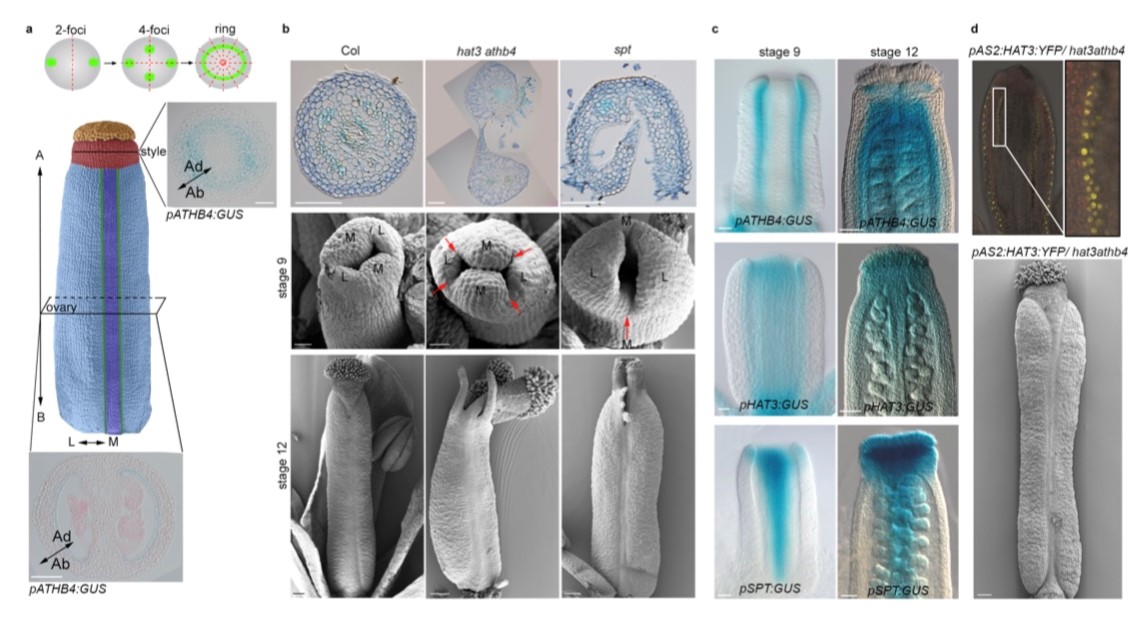
Coordination of biradial-to-radial symmetry and tissue polarity by HD-ZIP II proteins
The female reproductive organ, the gynoecium, of the model plant Arabidopsis thaliana is, so far, the only known example studied at the molecular level of a developing structure that reprograms itself over time in order to achieve a transition from bilateral to radial symmetry.
In a study conducted at the IBPM-CNR in collaboration with the John Innes Center (UK) and CREA-GB (Rome), the role of the transcription factors HAT3 and ATHB4 in the transition from bilateral to radial symmetry during formation of the style, the apical part of the gynoecium, has been studied in the model plant Arabidopsis thaliana.
In previous studies, it had been shown that the change from bilateral to radial symmetry of the gynoecium occurs through various stages that can be described by a change in the distribution of the hormone auxin. In particular, in the apical part of the developing gynoecium, auxin allocation goes from a two foci distribution (bilateral symmetry) to one with 4 foci (biradial), and finally to a complete ring (radial). This process is cooperatively controlled by the transcription factors bHLH SPATULA (SPT), INDEHISCENT and HECATE1, 2 and 3. The related mutants, spatula and indehiscent are blocked at the bilateral two-foci stage, while the triple mutant hecate1, 2, 3 is blocked at the four foci stage. In the work just published in the journal Nature Communications, the role of HAT3 and ATHB4, two transcription factors known as direct targets of SPATULA, is described in the process that will lead to the formation of the radialized organ. In the work it is shown that the HAT3 and ATHB4 genes are also positively controlled by the HECATE genes in a cooperative way with SPATULA. Through genetic analysis, the use of inducible transcription factors and fluorescent markers, it was possible to establish that HAT3 and ATHB4 control the distribution of auxin and sensitivity to the hormone cytokinin. A negative feedback control of HAT3 and ATHB4 on the SPATULA gene was also observed. This regulation is most likely responsible for the correct balance of auxin and cytokinin hormone signaling over time and space, so as to allow the transition of symmetry and the formation of the correct radialized organ.
With this work, we want to remember our colleague Ida Ruberti, who sadly passed away during the final phase of this work, for her fundamental role in the conceptualization of the project and the in-depth discussions on the results.

07/07/2021
Aspettando la Notte Europea Dei Ricercatori Talenti per la Scienza - 7, 9 E 10 Luglio 2021

Documenti:
NET_TALENTI-PER-LA-SCIENZA.pdf
Mappa TALENTI-.pdf
06/07/2021
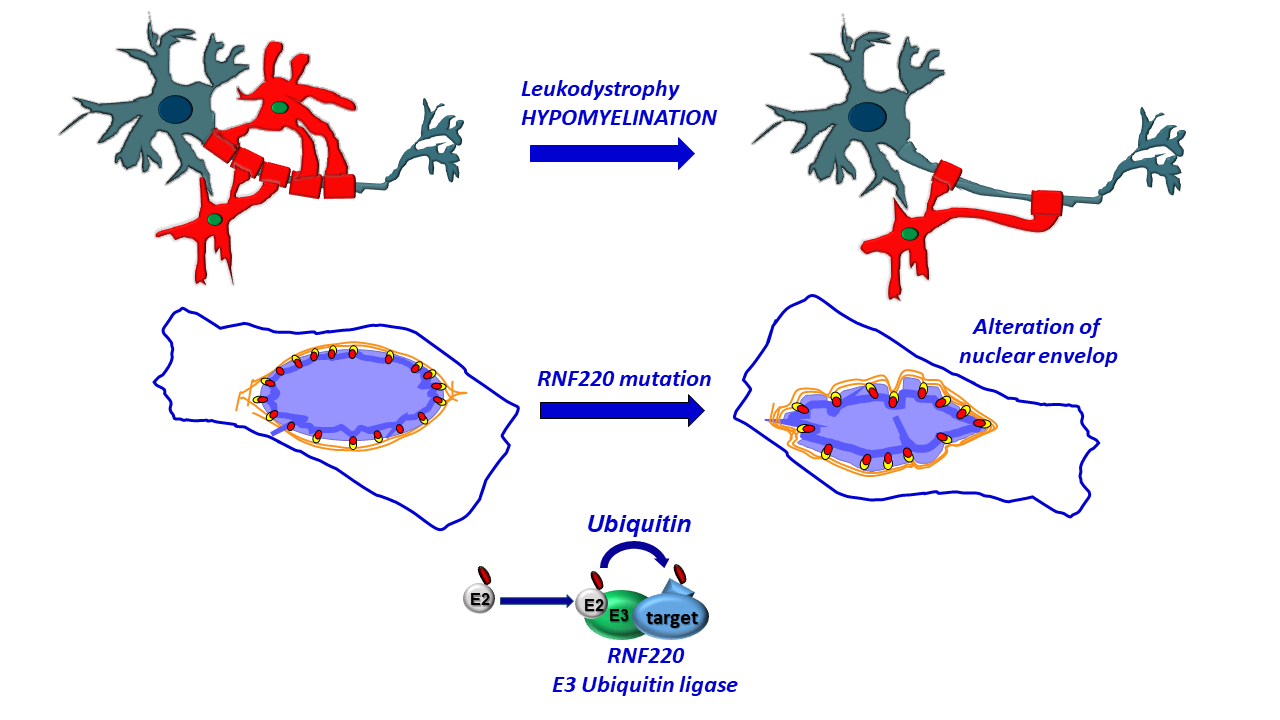
Biallelic mutations in RNF220 cause laminopathies featuring leukodystrophy, ataxia and deafness
We exploit Drosophila melanogaster to dissect the molecular mechanisms underlying neurologic and neurodegenerative disorders, and we are studying a novel form of leukodystrophy, linked to ataxia and sensorineural deafness.
Whole exome sequencing performed in several large consanguineous families, by our collaborators, allowed to identify two missense homozygous mutations that affect conserved residues of the ubiquitin E3 ligase RNF220, as causative events of a novel form of leukodystrophy. Mass spectrometry analysis identified lamin B1 as RNF220 binding partner. We demonstrated that RNF220 silencing in Drosophila specifically affects subcellular localization of fly lamin B1 orthologue, and promotes its aggregation. Further, RNF220 loss of function promotes significant neurodegenerative phenotypes in fly tissues, strongly supporting a functional connection between RNF220 and lamin B1.
Thus, we demonstrate that RNF220 is implicated in a novel form of leukodystrophy characterized by ataxia and sensorineural hearing loss, and show its role in the regulation of the nuclear lamina. Furthermore, we establish an in vivo fly disease model that could be exploited to design new therapeutic strategies.
03/06/2021
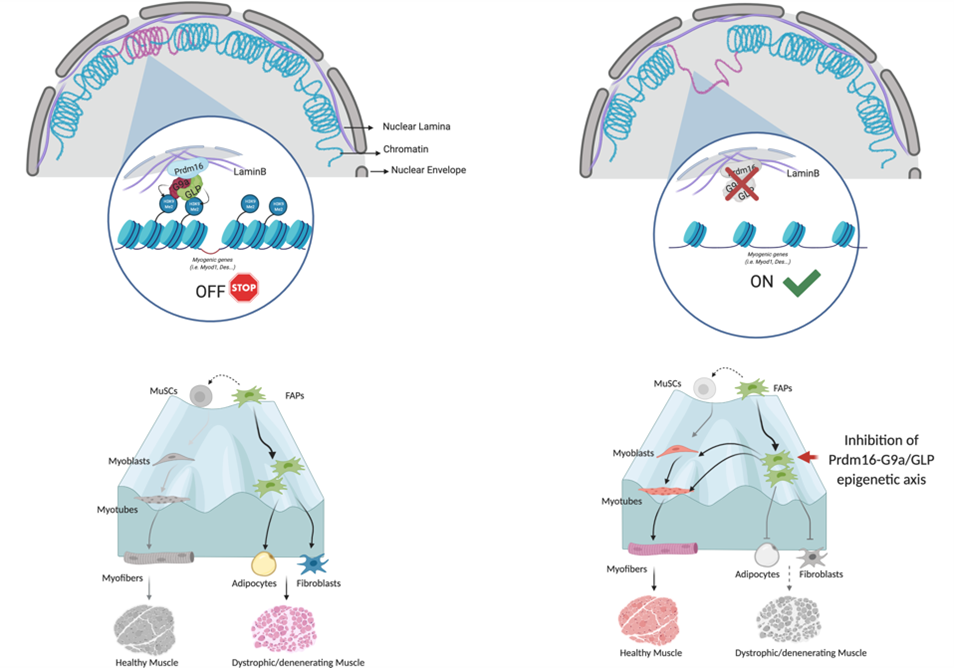
Prdm16-mediated H3K9 methylation controls fibro-adipogenic progenitors identity during skeletal muscle repair
Biferali et. al. (DOI: 10.1126/sciadv.abd9371)
In this work, we studied a mesenchymal stromal cell population (FIbro-Adipogenic Progenitros, FAPs) that plays a dual role in skeletal muscle homeostasis and regeneration. While they promote muscle regeneration by supporting paracrinally muscle stem cells. On the other hand having the potential to give rise to fibrotic and fat cells, they directly contribute to the degeneration of diseased muscles, such as those of patients suffering from Duchenne muscular dystrophy.
In this study we have identified a molecular mechanism of gene repression carried out by the Prdm16 / G9a / GLP proteins that is crucial in maintaining silent the ability of FAPs to promote the formation of muscle tissue.
In particular, this work shows that Prdm16 plays a fundamental role in blocking the regions of DNA encoding the muscle potential of FAPs at the nuclear periphery, recruiting the enzymes G9a and GLP to keep them silent. This discovery has an important translation and therapeutic potential. Indeed, the results of this work show that by inhibiting these proteins in mouse models of regeneration and dystrophy, it is possible to change the fate of these cells to consequently promote the regeneration of dystrophic muscles and, at the same time, slowing down their degeneration. In this work, we studied a mesenchymal stromal cell population (FIbro-Adipogenic Progenitros, FAPs) that plays a dual role in skeletal muscle homeostasis and regeneration. While they promote muscle regeneration by supporting paracrinally muscle stem cells. On the other hand having the potential to give rise to fibrotic and fat cells, they directly contribute to the degeneration of diseased muscles, such as those of patients suffering from Duchenne muscular dystrophy.
In this study we have identified a molecular mechanism of gene repression carried out by the Prdm16 / G9a / GLP proteins that is crucial in maintaining silent the ability of FAPs to promote the formation of muscle tissue.
In particular, this work shows that Prdm16 plays a fundamental role in blocking the regions of DNA encoding the muscle potential of FAPs at the nuclear periphery, recruiting the enzymes G9a and GLP to keep them silent. This discovery has an important translation and therapeutic potential. Indeed, the results of this work show that by inhibiting these proteins in mouse models of regeneration and dystrophy, it is possible to change the fate of these cells to consequently promote the regeneration of dystrophic muscles and, at the same time, slowing down their degeneration.
IBPM reference person: Chiara Mozzetta
Link: https://advances.sciencemag.org/content/7/23/eabd9371

04/03/2021
Cell Reports - Endothelin-1 drives invadopodia and interaction with mesothelial cells through ILK
DOI:https://doi.org/10.1016/j.celrep.2021.108800
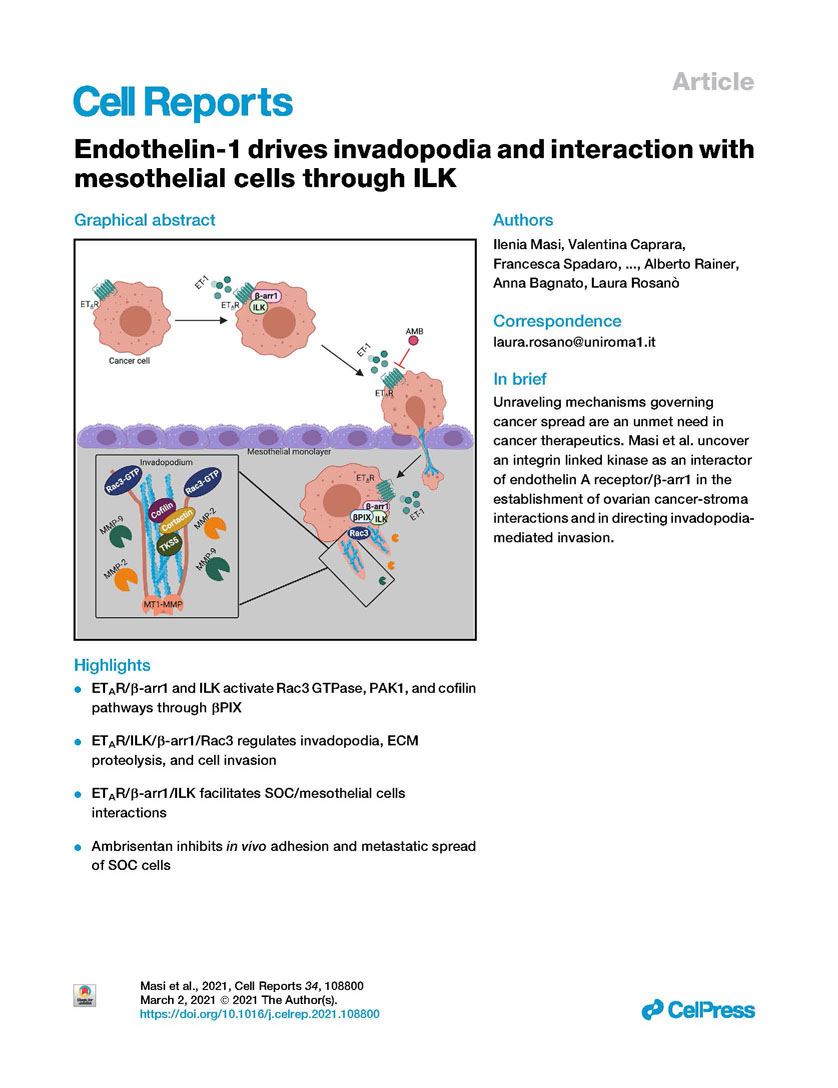
Documents: Endothelin-1 drives invadopodia and interaction with mesothelial cells through ILK.pdf
01/02/2021
Among the winners of Start Cup Lazio 2020 there is the CNR team composed by Dr Patrizia Brunetti (IBPM-CNR), Dr Davide Marzi (IBPM-CNR) and Dr Giancarlo Daniele (EGATO-VT1),
with the project “Hydro Fern” that was awarded with the Special Mention "Social Innovation", the Special Lazio Region Award,
the 4th place and the access to the Premio Nazionale dell’Innovazione (PNI 2020).

Documents: 5_Green-Clean-Biotech_LAZIO7_Avvenire_24_01_2021_pag3.pdf
Links:
https://www.cnr.it/it/news/9768/una-green-revolution-del-cnr-tra-i-vincitori-della-startcup-lazio-2020
https://www.tg24.info/regione-start-cup-lazio-2020-i-vincitori-della-6a-edizione/
https://www.repubblica.it/economia/rapporti/impresa-italia/mercati/2020/10/27/news/ecco_i_vincitori_della_6a_edizione_di_start_cup_lazio_2020-272047189/
28/01/2021
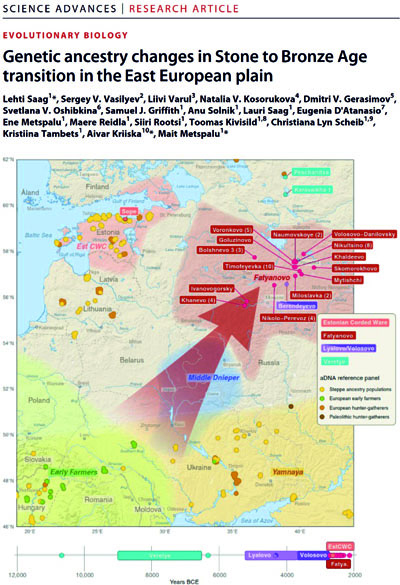
In this study, we produced genome-wide sequences from 30 ancient individuals from western Russia, including 3 Stone Age hunter-gatherers (10,800 to 4250 BCE) and 26 farmers from the Bronze Age Corded Ware complex Fatyanovo Culture (2900 to 2050 BCE). The analyses of the ancient DNA data allow us to show that Eastern European hunter-gatherer ancestry was present in Russia already from 10,000 BCE. What is more, a change in the ancestry is observed with the arrival of farming in the area, with the Fatyanovo samples showing both Steppe and Early farmer ancestry, similarly to other Corded Ware groups. Finally, we were also able to assess the genotypes at 113 phenotype-informative markers, showing allele frequency changes in markers related to pigmentation and diet between hunter-gatherers and Fatyanovo farmers. In conclusion, our data seems to suggest that the Fatyanovo-associated people likely originate from a fast migration toward the northeast from a region near Ukraine, where Steppe and Early farmer ancestries coexisted from around 3000 BCE.
IBPM reference person: Eugenia D'Atanasio
15/12/2020

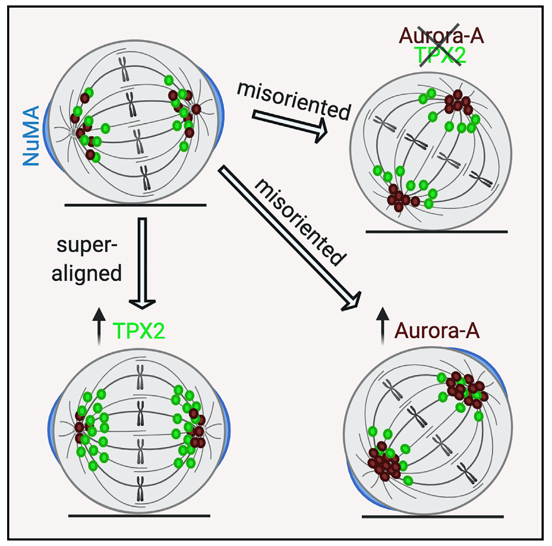 In this study we characterised the role of the Aurora-A kinase and its major regulator TPX2 in the control of mitotic spindle orientation in mammalian cells, a process which determines the position and fate of daughter cells. We highlighted a key role of TPX2 in the Aurora-A-mediated regulation of the orientation complex protein NuMA. We also describe opposite, microtubule-dependent, effects following increased Aurora-A or TPX2 levels in non-transformed cells, yielding-respectively- misoriented or “superaligned” spindles. Since high levels of Aurora-A and TPX2 are frequently observed in tumors, this study provides novel information to understand their pro-tumorigenic functions. It also opens the way to the investigation of the roles of the two proteins in other pathological contexts, particularly developmental or neurological disorders associated to defective cell division orientation.
In this study we characterised the role of the Aurora-A kinase and its major regulator TPX2 in the control of mitotic spindle orientation in mammalian cells, a process which determines the position and fate of daughter cells. We highlighted a key role of TPX2 in the Aurora-A-mediated regulation of the orientation complex protein NuMA. We also describe opposite, microtubule-dependent, effects following increased Aurora-A or TPX2 levels in non-transformed cells, yielding-respectively- misoriented or “superaligned” spindles. Since high levels of Aurora-A and TPX2 are frequently observed in tumors, this study provides novel information to understand their pro-tumorigenic functions. It also opens the way to the investigation of the roles of the two proteins in other pathological contexts, particularly developmental or neurological disorders associated to defective cell division orientation.
Work was collaboratively carried out by groups at the Institute of Molecular Biology and Pathology (IBPM) and at the European Institute of Oncology (IEO). Research groups at Sapienza University and at the Max Planck Institute of Molecular Physiology (Dortmund, DE) participated to the project.
IBPM reference person: Giulia Guarguaglini
17/8/2020
Link at https://rdcu.be/b6jAd
www.immunophenotype.org
twitter: immunosurveillance@TheCrick
This study identified a “core” immune signature in the peripheral blood of COVID-19 patients. Blood T cells were typically characterized by co-existing lymphopenia and highly activated/proliferative phenotype, especially in severe patients. Some signature traits, including basophil and plasmacytoid dendritic cell depletion, and T cell-cycling, correlated with disease severity. Notably, a triad of molecules — IP-10, interleukin-10 and interleukin-6— anticipated disease worsening.
The study, called COVID-IP, was led by prof. Adrian Hayday (Francis Crick Institute, King’s College London, London, UK). A cohort of 63 hospitalized COVID-19 patients at Guy’s and St Thomas’ Hospitals (London, UK) was studied. Dr. Francesca Di Rosa (Institute of Molecular Biology and Pathology, CNR, Rome, Italy) was a member of the international team. She contributed to the novelty by introducing the innovative T cell assay for cell cycle analysis, that was used for the first time in COVID-19 patients in this study.
13/7/2020
The identification in living cells of the protein FUS, crucially involved in Amyotrophic Lateral Sclerosis (ALS), as a direct interactor of HOTAIRM1 in motoneurons, together with the presence of a cytoplasmic isoform of HOTAIRM1 in motoneuron soma and neurites (yellow arrows), open the way for investigating a possible implication of this noncoding transcript in neurodegeneration.
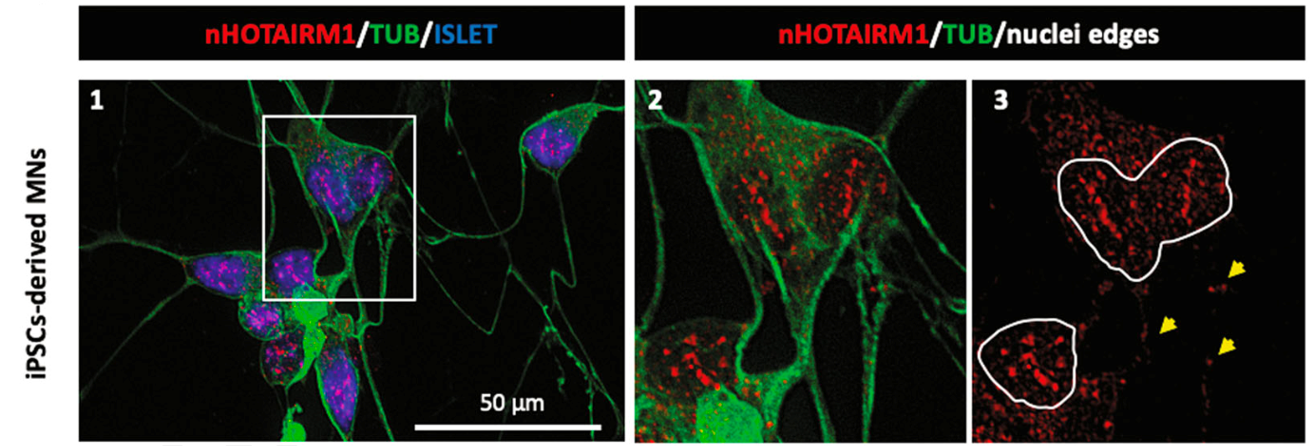
IBPM reference researchers: Pietro Laneve and Elisa Caffarelli
6/12/2019
IBPM at the Book Exhibition "Più libri più liberi": event on the DNA - WATCH THE VIDEO
Documents: PLPL_2019_PROGRAMMA_CNR.pdf
07/11/2019
In the school-work alternation project led by Maura Cardarelli and Patrizia Brunetti at IBPM, students of the Pascal High School train to use "green" technologies based on plant molecular biology to decontaminate soil from arsenic

Source : Rassegna Stampa CNR
08/10/2019
The 2019 Nobel prize to Physiology or Medicine for the discovery of the ability of cells to adapt to oxygen levels
Comment by Alessandro Giuffrè
 Source: https://www.cnr.it/rassegnastampa/19-10/191008/AJ1YUC.tif
Source: https://www.cnr.it/rassegnastampa/19-10/191008/AJ1YUC.tif
23/07/2019
Astrobiology. ASI, the Italian Space Agency, supports the project "LIFE IN SPACE", including the IBPM Unit coordinated by Giovanna Costanzo. On PLATINUM https://platinum-online.com/, the new magazine distributed by Il Sole 24 ore

10/07/2019
Documents: Meeting with delegation from Ikiam University.pdf
Meet in Italy for Life Sciences 2019
28/03/2019

This study highlights a novel role of the metabolic enzyme serine hydroxymethyl transferase (SHMT), in supporting the high proliferative capacity of cancer cells.
https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gkz129/5365575
12/03/2019
13/02/2019
16/01/2019
During mitotic division genomic DNA is equally partitioned between the daughter cells, and several cellular machineries are reorganized to ensure faithful chromosome segregation. For example, cellular processes such as transcription and mRNA processing (“splicing”) are shut down during mitosis. The work by Pellacani et al. shows that two splicing factors, Sf3A2 e Prp31, are not inactive during mitosis but directly mediate proper kinetochore- microtubule interactions. These findings open the way to future research aimed at identifying new “moonlighting proteins” involved in both nuclear processes occurring during interphase and regulation of mitosis. An accurate understanding of mitosis at the molecular level is crucial to elucidate the mechanisms of tumorigenesis and develop new anti-cancer therapies.
DUCHENNE DISTROPHY: ENGINEERING A NEW ARTIFICIAL GENE FOR THERAPEUTIC PURPOSES