Fax :
alice.pajoro@cnr.it
Researcher
National Research Council
Istitute of Molecular Biology and Pathology CNR
Sapienza University of Rome - Piazzale Aldo Moro 5, 00185 - Rome, Italy - Department of Biology and Biotechnologies C.Darwin - Building CU026 (General Physiology, ex BCS)
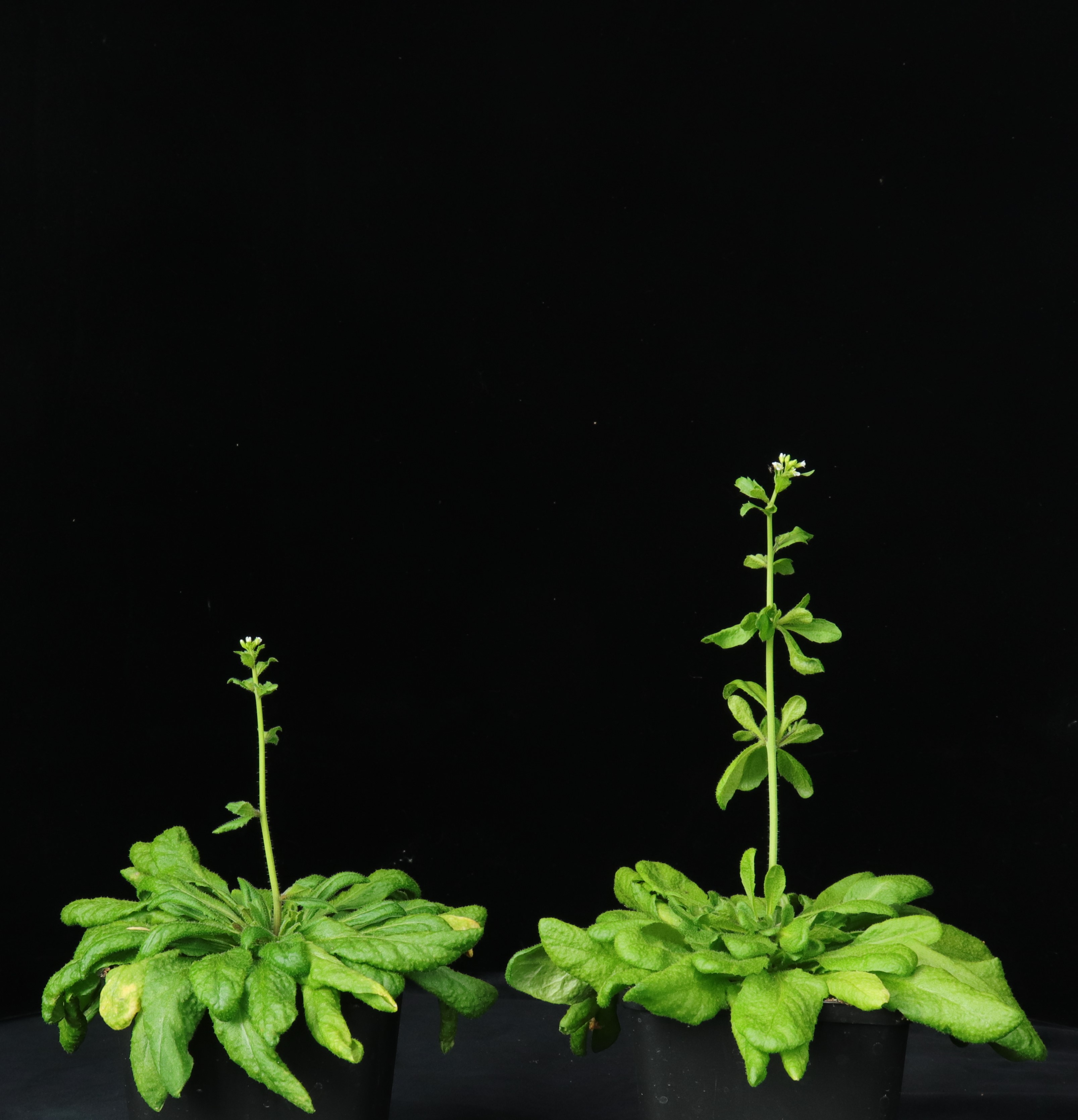
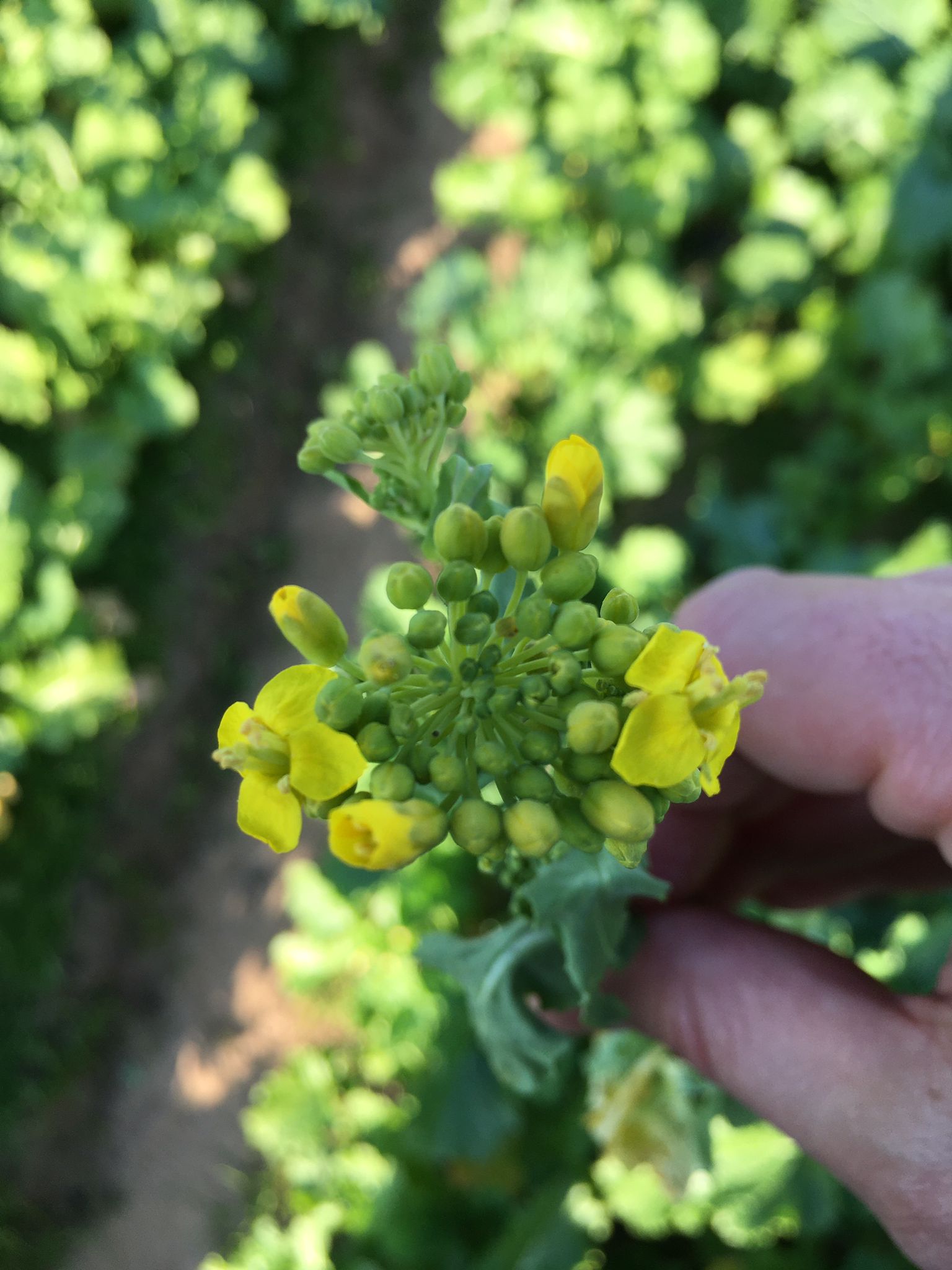
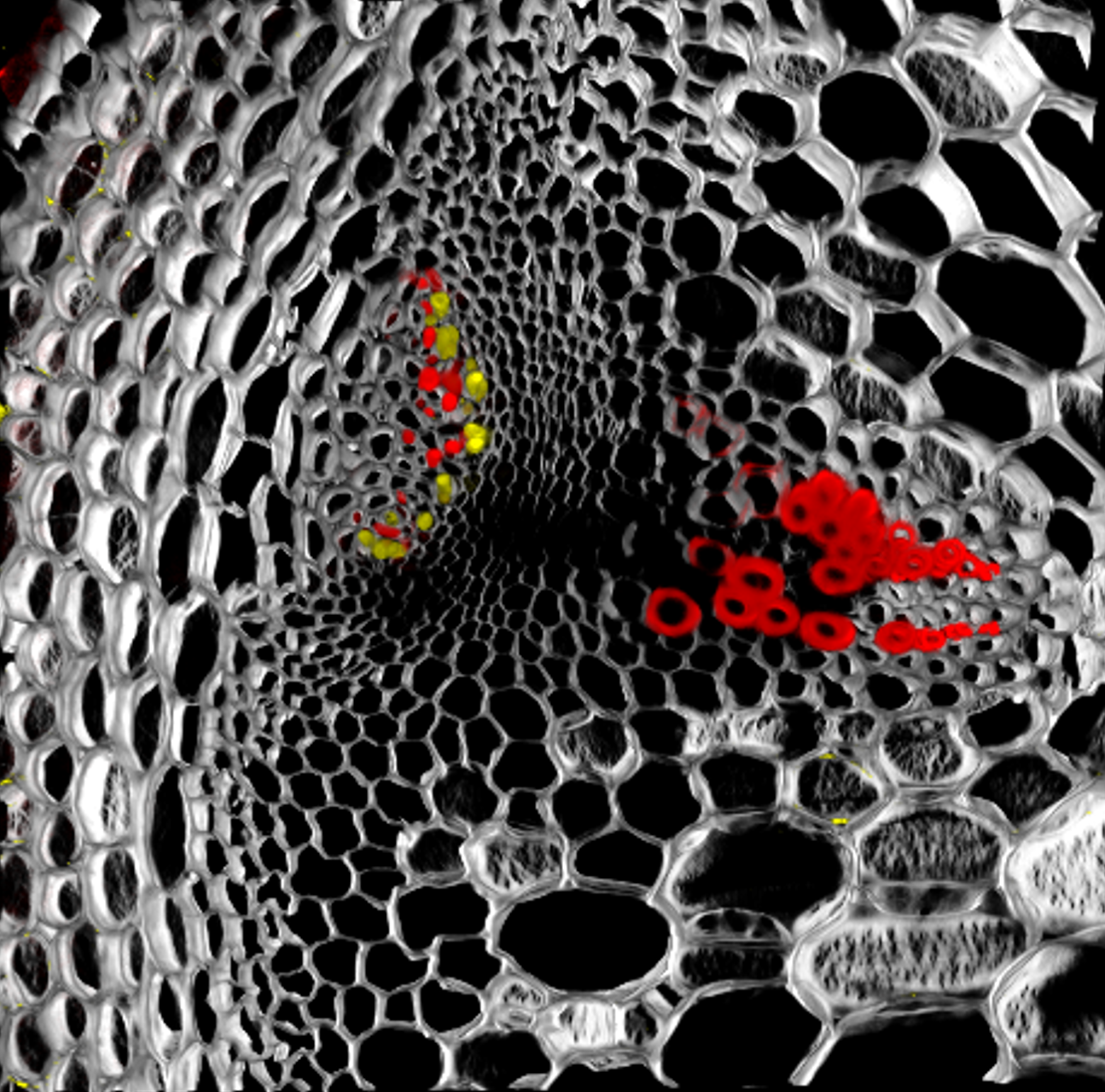
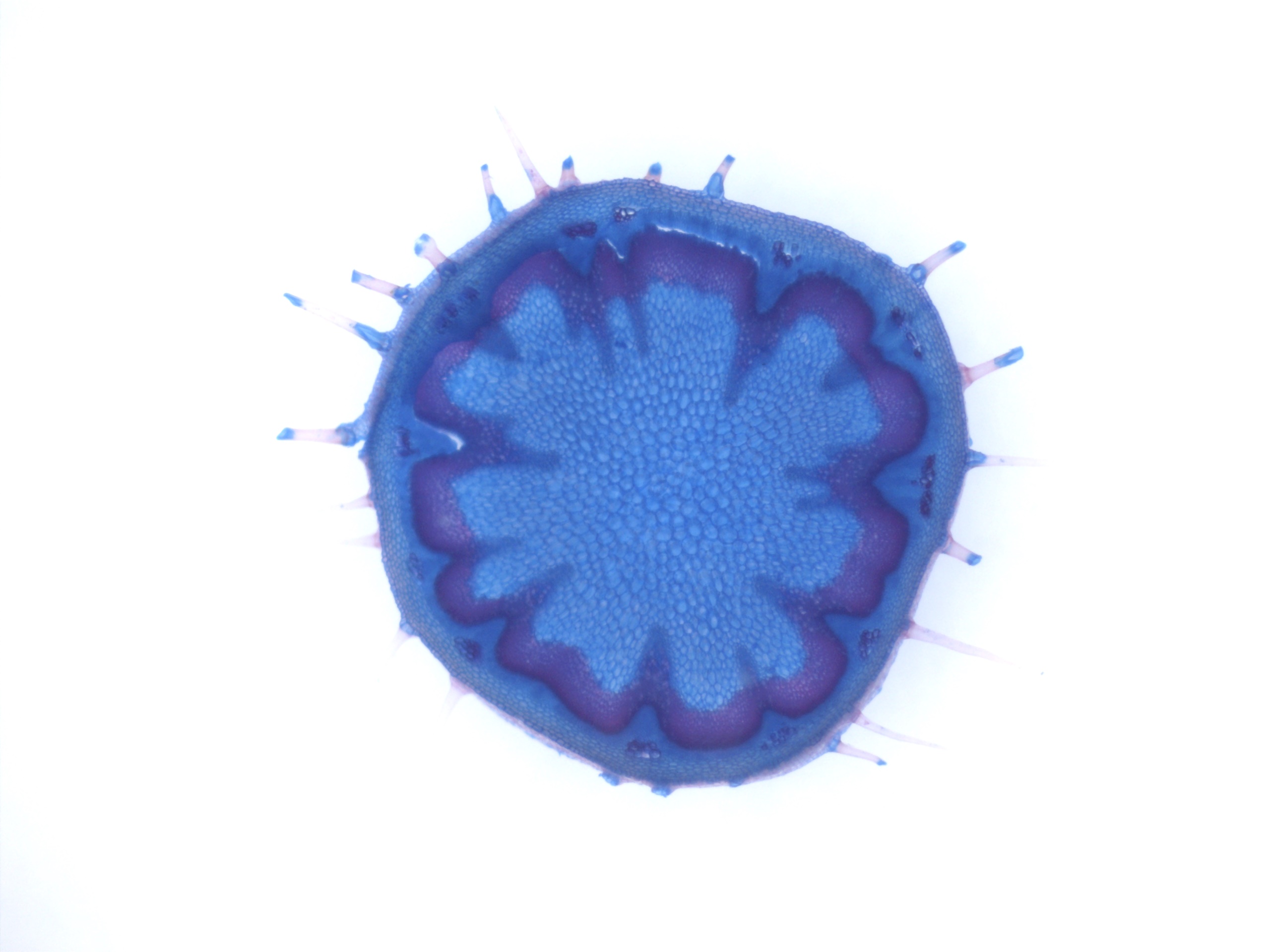
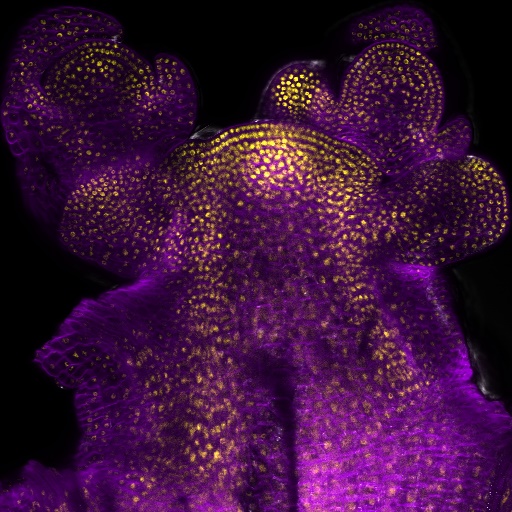
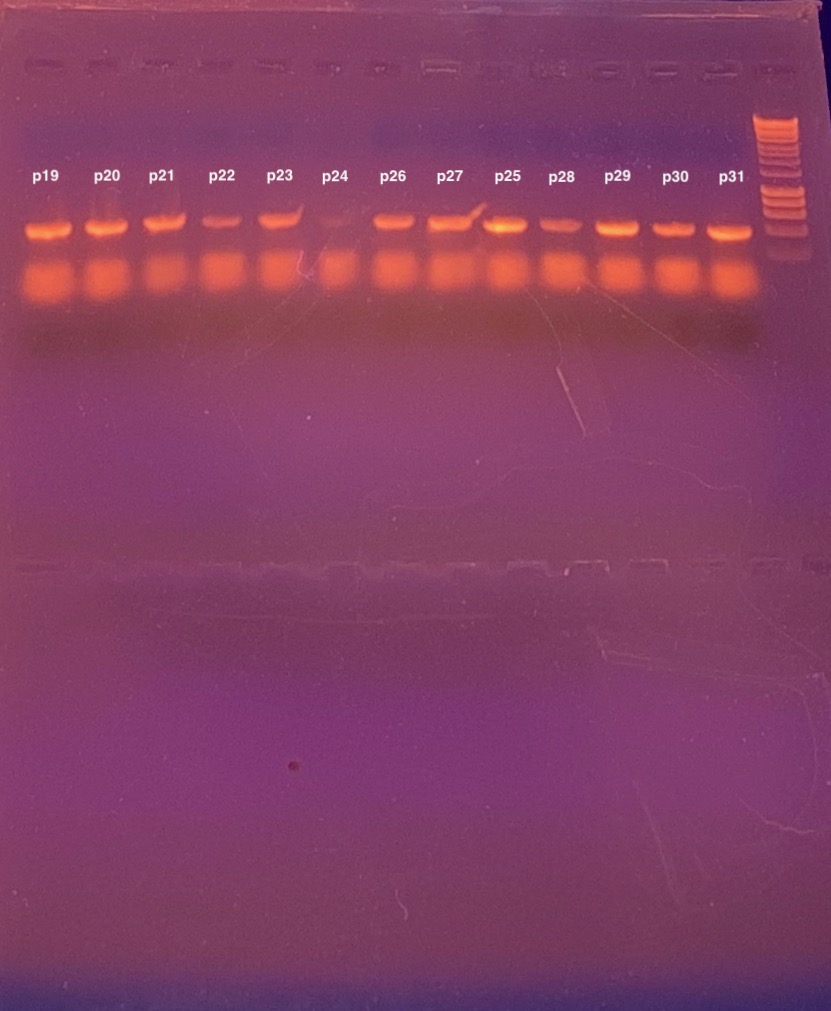
| My research projects focus on molecular mechanisms that regulate plant development in response to endogenous and environmental signals during the reproductive phase. Current projects: StemFlow: In plants, the transition from vegetative to reproductive growth requires coordinated developmental changes in different tissues. How the initiation of flower development at the apex of the plant is coordinated with alterations in stem growth to support the inflorescence is poorly understood. Key regulators of floral transition exhibit precise cellular expression patterns in stems, in the project we study how their expression patterns in stems and shoot apices are coordinated and define their functions in controlling stem growth. Funded by the Max-Planck Partnership. In collaboration with Prof. G. Coupland at Max-Planck Institute for Plant Breeding Research, Cologne, Gemany Top of the Crops: INTERAZIONI TRA LUCE E ORMONI NEL CONTROLLO DELL’APERTURA FIORALE IN SPECIE MODELLO E ORTIVE. L’apertura del fiore rappresenta l’ultimo stadio dello sviluppo fiorale ed è controllato da fattori ormonali che ambientali, la cui interazione definisce il successo e la durata del processo. Questo tratto impatta su molte colture ortive Brassicaceae, tra cui una tipica coltura laziale, la cima di rapa, in quanto una prematura apertura del fiore ne comporta un deprezzamento. Il progetto si articola su due fasi: l’identificazione dei geni responsabili dell’apertura dei fiori nella specie modello Arabidopsis thaliana che è anch’essa una Brassicacea e il successivo trasferimento della conoscenza alla coltura ortiva cime di rapa. Funded by Regione Lazio. In collaboration with Prof. G. Serino Dipartimento di Biologia e Biotecnologie C. Darwin, La Sapienza, & Dr D. Giannino, Istituto dei Sistemi Biologici, CNR, Roma, Italy EcoRap: Valorizzazione della collezione del germoplasma di cime di rapa pugliesi in un’ottica di economia sostenibile. Il progetto propone di agire su due fronti per rendere la coltura della cima di rapa più produttiva e sostenibile, da un lato individuare con approccio genomico le varianti strutturali della biodiversità delle cime associate al ritardo nell’antesi, dall’altro esplorare l’uso dei prodotti di scarto come fonte di composti micostatici e/o micotossici nei confronti di funghi. Nell’ottica di un’economia circolare i residui della coltura potranno essere convertiti in prodotto ad alto valore aggiunto, contribuendo così alla riduzione dei rifiuti e ai costi di smaltimento a carico degli agricoltori. Funded by Progetto Bio-Memory, Dipartimento di Scienze Bio-Agroalimentari In collaboration with Dr. Alessandra Ricelli, CNR-IBPM Functional Genomics Approaches for Improving Plant Inflorescence Architecture: Sufficient food production is threatened by the fast-growing world population and global climate change. To produce sufficient food for future generations in a sustainable manner, it is necessary to develop innovative strategies for plant production. This can only be achieved through an increased fundamental knowledge on how plants develop. This project intends to contribute to increase plant yield by studying pathways that control inflorescence development. Funded by MUR, PRIN2020. In collaboration with Prof M. Kater, Universita' degli Studi di Milano, Italy Rucol-ITA: Rucola is a food with high nutritional value because is reach of micronutrients and antioxidants. This characteristic has increased the consumer request of rucola in the European Market. Italy represent one of the major producers of Rucola in Europe. The project aims to select of new rucola varieties with higher performance. In collaboration with Prof. L. Faino at Dipartimento di Biologia Ambientale, La Sapienza, Roma, Italy |
|
| Dr. Maura Cardarelli, CNR-IBPM Prof. G. Coupland at Max-Planck Institute for Plant Breeding Research, Cologne, Gemany Prof. G. Serino Dipartimento di Biologia e Biotecnologie C. Darwin, La Sapienza Dr D. Giannino, Istituto dei Sistemi Biologici, CNR, Roma, Italy Prof M. Kater, Universita' degli Studi di Milano, Italy Prof. L. Faino at Dipartimento di Biologia Ambientale, La Sapienza, Roma, Italy Prof. R. Immink, Wageningen University, The Netherlands |
2019-present: Researcher/Junior PI Istituto di Biologia a patologia molecolari (IBPM) Consiglio Nazionale delle Ricerche c/o Dipartimento di Biologia e Biotecnologie “Charles Darwin”. 2014 – 2017 Post Doc in Prof R. Immink’s group, Plant Research International, Wageningen, The Netherlands. The research focuses on understand the impact of ambient temperature in regulation of developmental processes in Arabidopsis, within the ERA-CAPS network, FLOWPLAST. 2009 – 2014 PhD in Molecular Biology in Prof. G. Angenent’s department supervised by Dr. K. Kaufmann, Wageningen University, Wageningen, The Netherlands. My project was carried out in the framework of the Marie Curie ITN project SYSFLO. The project focused on the gene regulatory networks underlying flower development in Arabidopsis. 2008 – 2009 Intern at the Universitá degli Studi di Milano in Prof. M. Kater’s group. My project focused on the role of OsMADS13 and OsMADS21 in rice ovule and kernel development and the functional conservation of gene function between Arabidopsis and rice. 2006 Intern at the Universitá degli Studi di Milano in Prof. G. Gavazzi and Prof G. Consonni’s group. The project focused on the characterization fused leaves mutant (fdl) in Zea mais” |