Fax :
patrizia.filetici@uniroma1.it
Senior Researcher
Istitute of Molecular Biology and Pathology - National Research Council
Department of Biology and Biotecnology - Charles Darwin
Sapienza University of Rome - Istituto Fisiologia Generale I piano, stanza 5 e 14 P.Le A.Moro 5 00185 Roma
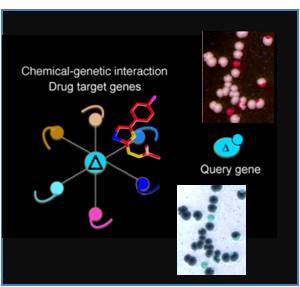
An epigenetic trait is a stably heritable phenotype resulting from changes in a chromosome without alterations in the DNA sequence C. Waddington 1950s Epigenetics regulates the genome through dynamic waves of chromatin condensation and decondensation through pleiotropic key mechanisms controling several interlaced pathways. We use simple Yeast as awesome model to study higher Eukaryotes and human cells
Epigenetic Regulators, Post-Translational Modifications and Their Cross-Talk, Studies From Model Organisms to Human cells and Cancer
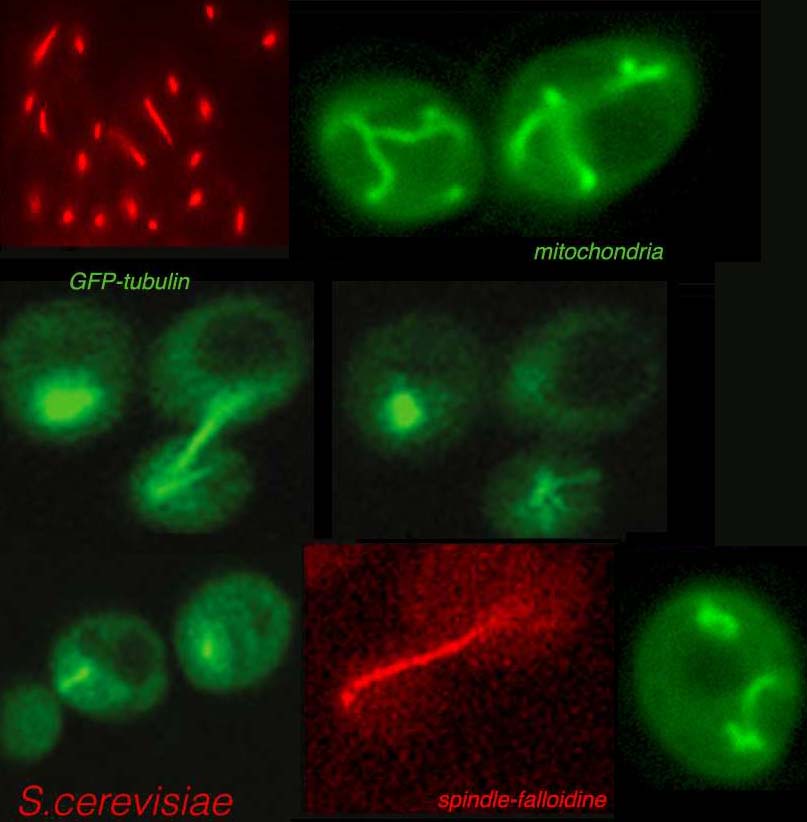
Post-translational modifications mark histone tails to modulate the accessibility of chromatin that affects the pattern of gene expression and cell differentiation. Our laboratory is interested in the comprehension of epigenetic mechanisms and role of Post-Translational Modifications such as Acetylation regulating the genome accessibility and modification of non histone proteins. Among research achievements obtained in the group there were the first studies on the chromatin remodeling activity of HAT Gcn5 and the discovery of the role of the chromatin Bromodomain in reading the Ac-Lys of histone tails. We have gained other results in the study of the pleiotropic multifunctional role of the HAT Gcn5, reporting its function not only as coactivator of gene transcription and in chromatin organization but also as regulator of cell cycle progression and its specific role in the organization and epigenetic mark of the centromere-kinetochore region. Recently we are interested in elucidating novel aspects of the acetylation dependent regulation of respiratorymetabolism in order to determine the role of acetylation on mitochondria functions, cell mitophagy and respiratory functions.
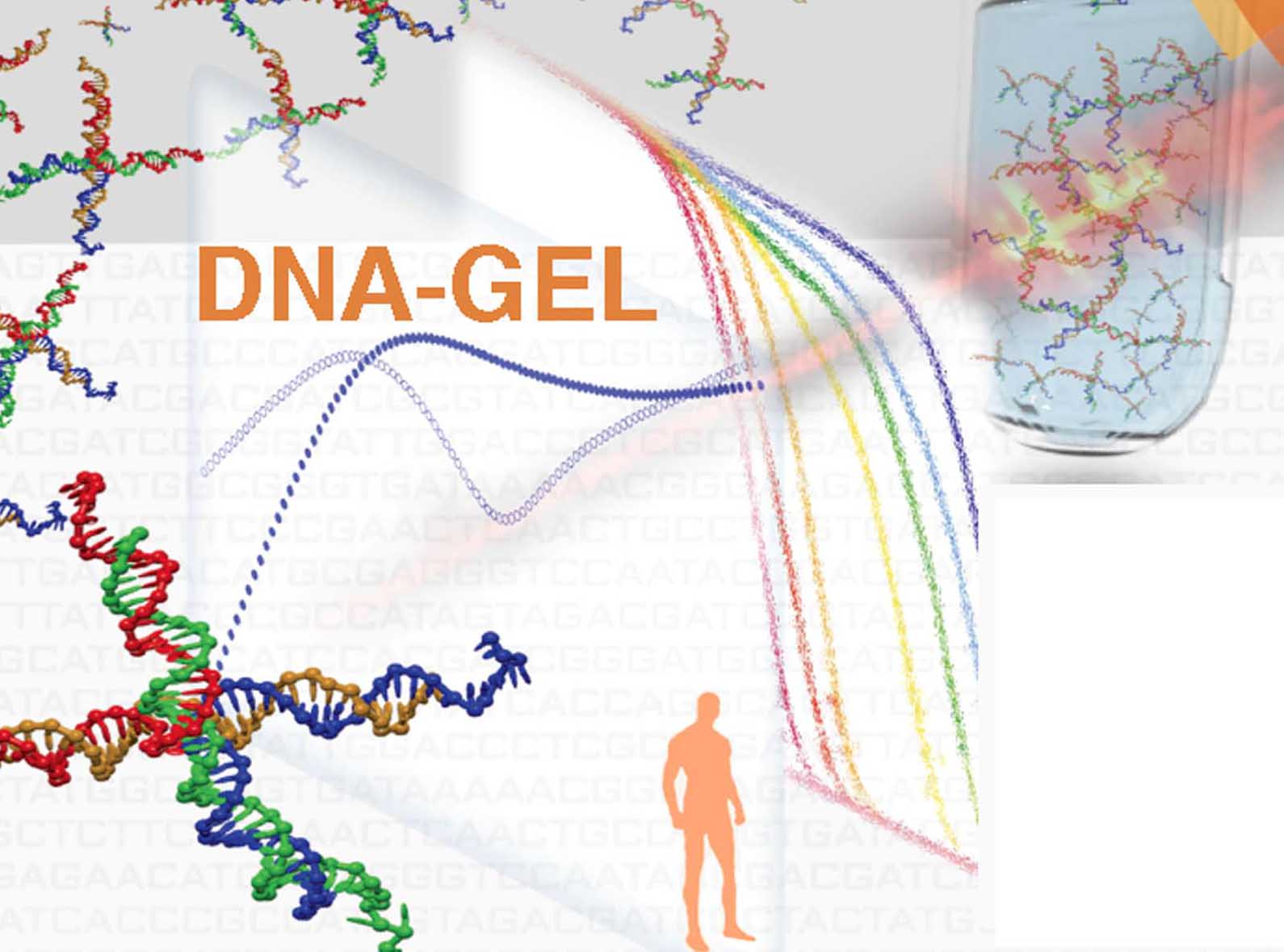
A novel area of research in addition is involved in the design and utilization of biocompatibile DNA-hydrogels with atypical behaviour . This new project will be developed for the achievement of producing novel bio-compatible nanomaterial for the delivery of bioactive molecules in human tissues. |
Montanari A, Leo M, De Luca V, Filetici P, Francisci S. 2019 Gcn5 histone acetyltransferase is present in the mitoplasts. Biol Open. 8 (2). pii: bio041244. doi: 10.1242/bio.041244.
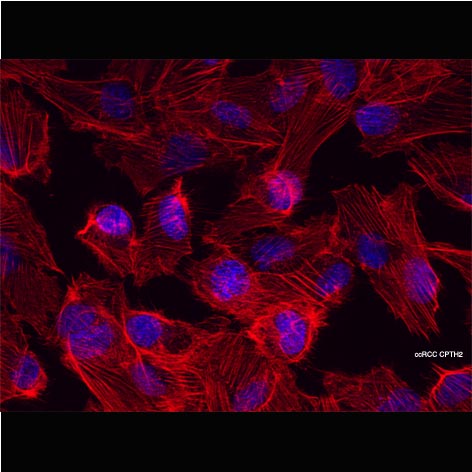
E. Cocco, M. Leo, C. Canzonetta, S. Di Vito, A. Mai, D. Rotili, A. Di Napoli, A. Vecchione, C. De Nunzio, P. Filetici and A. Stoppacciaro. 2018. KAT3B-p300 and H3AcK18/H3AcK14 levels are prognostic markers for kidney ccRCC tumor aggressiveness and target of KAT inhibitor CPTH2. Clinical Epigenetics. 10:44 https://doi.org/10.1186/s13148-018-0473-4
B. Ricci, S. Monte, S. Vernarecci, C. Canzonetta, P. Ballario, P. Filetici. 2017. Growth arrest in yeast with aberrant kinetochore is rescued by the loss of Gcn5. Matters ISSN: 2297-8240 doi:10.19185/matters.201705000006
Canzonetta C, Leo M, Guarino SR, Montanari A, Francisci S, Filetici P. 2016 SAGA complex and Gcn5 are necessary for respiration in budding yeast.Biochim Biophys Acta. 1863(12):3160-3168.
Bomboi F, Romano F, Leo M, Fernandez-Castanon J, Cerbino R, Bellini T, Bordi F, Filetici P, Sciortino F. 2016 Re-entrant DNA gels. Nat Commun. 7:13191.
Canzonetta C, Vernarecci S, Iuliani M, Marracino C, Belloni C, Ballario P, Filetici P. 2015 SAGA DUB-Ubp8 Deubiquitylates Centromeric Histone Variant Cse4. G3 (Bethesda). 6(2):287-98.
Lenoci A, Tomassi S, Conte M, Benedetti R, Rodriguez V, Carradori S, Secci D, Castellano S, Sbardella G, Filetici P, Novellino E, Altucci L, Rotili D, Mai A. 2014 Quinoline-based p300 histone acetyltransferase inhibitors with pro-apoptotic activity in human leukemia U937 cells. ChemMedChem. 9 (3):542-8.
Brenna A, Grimaldi B, Filetici P, Ballario P. 2012 Physical association of the WC-1 photoreceptor and the histone acetyltransferase NGF-1 is required for blue light signal transduction in Neurospora crassa. Mol Biol Cell. 23(19):3863-72.
Trisciuoglio D, Ragazzoni Y, Pelosi A, Desideri M, Carradori S, Gabellini C, Maresca G, Nescatelli R, Secci D, Bolasco A, Bizzarri B, Cavaliere C, D'Agnano I, Filetici P, Ricci-Vitiani L, Rizzo MG, Del Bufalo D. 2012 CPTH6, a thiazole derivative, induces histone hypoacetylation and apoptosis in human leukemia cells. Clin Cancer Res. 18(2):475-86. doi: 10.1158/1078-0432.CCR-11-0579.
Vernarecci S, Tosi F, Filetici P. 2010 Tuning acetylated chromatin with HAT inhibitors: a novel tool for therapy. Epigenetics. 5(2):105-11.
Chimenti F, Bizzarri B, Maccioni E, Secci D, Bolasco A, Chimenti P, Fioravanti R, Granese A, Carradori S, Tosi F, Ballario P, Vernarecci S, Filetici P. 2009 A novel histone acetyltransferase inhibitor modulating Gcn5 network: cyclopentylidene-[4-(4'-chlorophenyl)thiazol-2-yl)hydrazone. J Med Chem. 52(2):530-6.
Vernarecci S, Ornaghi P, Bâgu A, Cundari E, Ballario P, Filetici P. 2008 Gcn5p plays an important role in centromere kinetochore function in budding yeast. Mol Cell Biol. 28(3):988-96.
Vernarecci S, Colotti G, Ornaghi P, Schiebel E, Chiancone E, Filetici P. 2007 The yeast penta-EF protein Pef1p is involved in cation-dependent budding and cell polarization. Mol Microbiol. 65(4):1122-38.
Mai A, Rotili D, Tarantino D, Ornaghi P, Tosi F, Vicidomini C, Sbardella G, Nebbioso A, Miceli M, Altucci L, Filetici P. 2006 Small-molecule inhibitors of histone acetyltransferase activity: identification and biological properties. J Med Chem. 49(23):6897-907.
Grimaldi B, Coiro P, Filetici P, Berge E, Dobosy JR, Freitag M, Selker EU, Ballario P. 2006 The Neurospora crassa White Collar-1 dependent blue light response requires acetylation of histone H3 lysine 14 by NGF-1. Mol Biol Cell. 17(10):4576-83.
Ishida C, Aranda C, Valenzuela L, Riego L, Deluna A, Recillas-Targa F, Filetici P, López-Revilla R, González A. 2006 The UGA3-GLT1 intergenic region constitutes a promoter whose bidirectional nature is determined by chromatin organization in Saccharomyces cerevisiae. Mol Microbiol. 59(6):1790-806.
Filetici P, P O, Ballario P. 2001 The bromodomain: a chromatin browser? Front Biosci. 6:D866-76.
Owen DJ, Ornaghi P, Yang JC, Lowe N, Evans PR, Ballario P, Neuhaus D, Filetici P, Travers AA. 2000 The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 19(22):6141-9.
Ornaghi P, Ballario P, Lena AM, González A, Filetici P. 1999 The bromodomain of Gcn5p interacts in vitro with specific residues in the N terminus of histone H4. J Mol Biol. 287(1):1-7.
Filetici P, Aranda C, Gonzàlez A, Ballario P. 1998 GCN5, a yeast transcriptional coactivator, induces chromatin reconfiguration of HIS3 promoter in vivo. Biochem Biophys Res Commun. 242(1):84-7.
|
De Palma A, Fanelli G, Cretella E, De Luca V, Palladino RA, Panzeri V, Roffia V, Saliola M, Mauri P, Filetici P. Gcn5p and Ubp8p Affect Protein Ubiquitylation and Cell Proliferation by Altering the Fermentative/Respiratory Flux Balance in Saccharomyces cerevisiae. mBio. 2020 Aug 11;11(4):e01504-20. doi: 10.1128/mBio.01504-20. PMID: 32788380
Enrico Lattuada, Manuela Leo, Debora Caprara, Luisa Salvatori, Antonella Stoppacciaro, Francesco Sciortino, Patrizia Filetici DNA-GEL, Novel Nanomaterial for Biomedical Applications and Delivery of Bioactive Molecules Front. Pharmacol., 04 September 2020, 11- 1345 | https://doi.org/10.3389/fphar.2020.01345 |
Montanari A, Leo M, De Luca V, Filetici P, Francisci S. 2019 Gcn5 histone acetyltransferase is present in the mitoplasts. Biol Open. 8 (2). pii: bio041244. doi: 10.1242/bio.041244.
E. Cocco, M. Leo, C. Canzonetta, S. Di Vito, A. Mai, D. Rotili, A. Di Napoli, A. Vecchione, C. De Nunzio, P. Filetici and A. Stoppacciaro. 2018. KAT3B-p300 and H3AcK18/H3AcK14 levels are prognostic markers for kidney ccRCC tumor aggressiveness and target of KAT inhibitor CPTH2. Clinical Epigenetics. 10:44 https://doi.org/10.1186/s13148-018-0473-4
B. Ricci, S. Monte, S. Vernarecci, C. Canzonetta, P. Ballario, P. Filetici. 2017. Growth arrest in yeast with aberrant kinetochore is rescued by the loss of Gcn5. Matters ISSN: 2297-8240 doi:10.19185/matters.201705000006
Canzonetta C, Leo M, Guarino SR, Montanari A, Francisci S, Filetici P. 2016 SAGA complex and Gcn5 are necessary for respiration in budding yeast.Biochim Biophys Acta. 1863(12):3160-3168.
Bomboi F, Romano F, Leo M, Fernandez-Castanon J, Cerbino R, Bellini T, Bordi F, Filetici P, Sciortino F. 2016 Re-entrant DNA gels. Nat Commun. 7:13191.
Canzonetta C, Vernarecci S, Iuliani M, Marracino C, Belloni C, Ballario P, Filetici P. 2015 SAGA DUB-Ubp8 Deubiquitylates Centromeric Histone Variant Cse4. G3 (Bethesda). 6(2):287-98.
Lenoci A, Tomassi S, Conte M, Benedetti R, Rodriguez V, Carradori S, Secci D, Castellano S, Sbardella G, Filetici P, Novellino E, Altucci L, Rotili D, Mai A. 2014 Quinoline-based p300 histone acetyltransferase inhibitors with pro-apoptotic activity in human leukemia U937 cells. ChemMedChem. 9 (3):542-8.
Brenna A, Grimaldi B, Filetici P, Ballario P. 2012 Physical association of the WC-1 photoreceptor and the histone acetyltransferase NGF-1 is required for blue light signal transduction in Neurospora crassa. Mol Biol Cell. 23(19):3863-72.
Trisciuoglio D, Ragazzoni Y, Pelosi A, Desideri M, Carradori S, Gabellini C, Maresca G, Nescatelli R, Secci D, Bolasco A, Bizzarri B, Cavaliere C, D'Agnano I, Filetici P, Ricci-Vitiani L, Rizzo MG, Del Bufalo D. 2012 CPTH6, a thiazole derivative, induces histone hypoacetylation and apoptosis in human leukemia cells. Clin Cancer Res. 18(2):475-86. doi: 10.1158/1078-0432.CCR-11-0579.
Vernarecci S, Tosi F, Filetici P. 2010 Tuning acetylated chromatin with HAT inhibitors: a novel tool for therapy. Epigenetics. 5(2):105-11.
Chimenti F, Bizzarri B, Maccioni E, Secci D, Bolasco A, Chimenti P, Fioravanti R, Granese A, Carradori S, Tosi F, Ballario P, Vernarecci S, Filetici P. 2009 A novel histone acetyltransferase inhibitor modulating Gcn5 network: cyclopentylidene-[4-(4'-chlorophenyl)thiazol-2-yl)hydrazone. J Med Chem. 52(2):530-6.
Vernarecci S, Ornaghi P, Bâgu A, Cundari E, Ballario P, Filetici P. 2008 Gcn5p plays an important role in centromere kinetochore function in budding yeast. Mol Cell Biol. 28(3):988-96.
Vernarecci S, Colotti G, Ornaghi P, Schiebel E, Chiancone E, Filetici P. 2007 The yeast penta-EF protein Pef1p is involved in cation-dependent budding and cell polarization. Mol Microbiol. 65(4):1122-38.
Mai A, Rotili D, Tarantino D, Ornaghi P, Tosi F, Vicidomini C, Sbardella G, Nebbioso A, Miceli M, Altucci L, Filetici P. 2006 Small-molecule inhibitors of histone acetyltransferase activity: identification and biological properties. J Med Chem. 49(23):6897-907.
Grimaldi B, Coiro P, Filetici P, Berge E, Dobosy JR, Freitag M, Selker EU, Ballario P. 2006 The Neurospora crassa White Collar-1 dependent blue light response requires acetylation of histone H3 lysine 14 by NGF-1. Mol Biol Cell. 17(10):4576-83.
Ishida C, Aranda C, Valenzuela L, Riego L, Deluna A, Recillas-Targa F, Filetici P, López-Revilla R, González A. 2006 The UGA3-GLT1 intergenic region constitutes a promoter whose bidirectional nature is determined by chromatin organization in Saccharomyces cerevisiae. Mol Microbiol. 59(6):1790-806.
Filetici P, P O, Ballario P. 2001 The bromodomain: a chromatin browser? Front Biosci. 6:D866-76.
Owen DJ, Ornaghi P, Yang JC, Lowe N, Evans PR, Ballario P, Neuhaus D, Filetici P, Travers AA. 2000 The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 19(22):6141-9.
Ornaghi P, Ballario P, Lena AM, González A, Filetici P. 1999 The bromodomain of Gcn5p interacts in vitro with specific residues in the N terminus of histone H4. J Mol Biol. 287(1):1-7.
Filetici P, Aranda C, Gonzàlez A, Ballario P. 1998 GCN5, a yeast transcriptional coactivator, induces chromatin reconfiguration of HIS3 promoter in vivo. Biochem Biophys Res Commun. 242(1):84-7.
|
De Palma A, Fanelli G, Cretella E, De Luca V, Palladino RA, Panzeri V, Roffia V, Saliola M, Mauri P, Filetici P. Gcn5p and Ubp8p Affect Protein Ubiquitylation and Cell Proliferation by Altering the Fermentative/Respiratory Flux Balance in Saccharomyces cerevisiae. mBio. 2020 Aug 11;11(4):e01504-20. doi: 10.1128/mBio.01504-20. PMID: 32788380
Enrico Lattuada, Manuela Leo, Debora Caprara, Luisa Salvatori, Antonella Stoppacciaro, Francesco Sciortino, Patrizia Filetici DNA-GEL, Novel Nanomaterial for Biomedical Applications and Delivery of Bioactive Molecules Front. Pharmacol., 04 September 2020, 11- 1345 | https://doi.org/10.3389/fphar.2020.01345 |
Research Group |
|
|