Fax :+39064440812
ernesto.dimauro@uniroma1.it
Senior Associate
Department of Biology and Biotecnology - Charles Darwin
Sapienza University of Rome
Istituto Fisiologia Generale - Primo piano P.Le A.Moro 5 00185 Roma
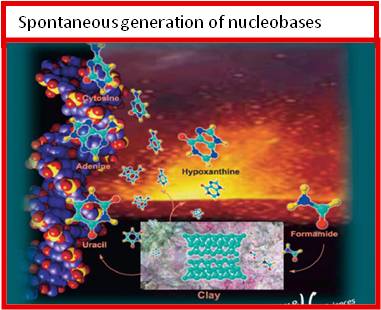
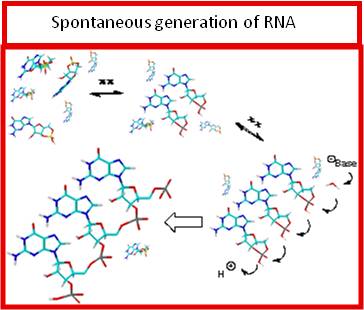
Life is made of the intimate interaction of metabolism and genetics, both built around the chemistry of the most common elements of the Universe (hydrogen, oxygen, nitrogen, carbon). The transmissible interaction of metabolic and genetic cycles results in the hypercycles of organization and de-organization of chemical information, of living and non-living. The origin-of-life quest has long been split in several attitudes exemplified by the aphorisms “genetics-first” or “metabolism-first”. Recently, the opposition between these approaches has been overstepped by more unitary theoretical and experimental frames taking into account energetic, evolutionary, proto-metabolic and ur-environmental aspects. Nevertheless, a unitary and simple chemical frame is still needed that could afford in a single “warm little pond” both the precursors of the synthetic pathways eventually leading to RNA and to the key components of the central metabolic cycles, possibly connected with the synthesis of fatty acids. In order to approach the problem of the origin of life it is therefore reasonable to start from the assumption that both metabolism and genetics had a common origin, shared a common chemical frame, were embedded in physical-chemical conditions favourable for the onset of both. Our research group has established the synthesis of nucleic bases, acyclonucleosides, nucleotides, biogenic carboxylic acids, sugars, amino sugars, aminoacids and condensing agents in a single and simple physical-chemical frame. Conditions for the spontaneous generation of RNA have been discovered. |
G. COSTANZO, R. SALADINO, G. BOTTA, AESSANDRA GIORGI, A. SCIPIONI, S. PINO and E. DI MAURO Generation of RNA molecules by base catalyzed click-like reaction CHEMBIOCHEM (2012) Mar 30. doi: 10.1002/cbic.201200068. [Epub ahead of print]
R. SALADINO, C. CRESTINI, S. PINO, G. COSTANZO, E. DI MAURO Formamide and the origin of life PHYSICS of LIFE REVIEWS (2012) 9, 84-104 doi:10.1016/j.plrev.2011.12.002
S. PINO, E. DI MAURO Gaia Universalis in “Consciousness and the Universe: Quantum Physics, Evolution, Brain & Mind” by R.Penrose , S. Hameroff, H. P. Stapp, D. Chopra, XIV. Consciousness and the Universe. JOURNAL OF COSMOLOGY Vol. 14, pp. 1251-1259, Cosmology Science Publishers, 2011
S. PINO, M. BIASIUCCI, M. SCARDAMAGLIA, G. GIGLI, MG. BETTI, C. MARIANI, E. DI MAURO Nonenzymatic Ligation of an RNA Oligonucleotide Analyzed by Atomic Force Microscopy. THE JOURNAL OF PHYSICAL CHEMISTRY B (2011) 115(19):6296-303
S. PINO, E. TRIFONOV and E. DI MAURO On the observable transition to living matter GENOMICS, PROTEOMICS AND BIOINFORMATICS, (2011) 9(1-2):7-14.
S. PINO, G. COSTANZO, A. GIORGI and E. DI MAURO Sequence Complementarity-Driven Nonenzymatic Ligation of RNA BIOCHEMISTRY, (2011) 50:2994-3003.
G. COSTANZO, S. PINO, F. CICIRIELLO, E. DI MAURO, Generation of long RNA chains in water THE JOURNAL OF BIOLOGICAL CHEMISTRY (2009) 284, 33206–33216
S. PINO, F. CICIRIELLO, G. COSTANZO, E. DI MAURO, Nonenzymatic RNA Ligation in Water THE JOURNAL OF BIOLOGICAL CHEMISTRY (2008) 283, 36494-36503
M.A. VEGA PALAS, S. VENDITTI and E. DI MAURO Telomeric transcriptional silencing in a natural context NATURE GENETICS 15, 232-233 (1997)
E. DI MAURO, L. SNIDER, P. MARINO, A. LAMBERTI, A. COPPO and G.P. TOCCHINI- VALENTINI Rifampicin sensitivity of the components of DNA dependent RNA polymerase NATURE 222, 533-537 (1969) |
http://bbcd.bio.uniroma1.it/bbcd/users/di-mauro-ernesto |
Born: 12.08.1945 Present position and title: Full Professor of Molecular Biology at “Dipartimento di Genetica e Biologia Molecolare” Università di Roma “Sapienza”, Rome Italy. 1967 Laurea in Molecular Biology, University of Rome, “Sapienza” 1967-1968 Intl. Laboratory of Genetics & Biophysics (Naples), Researcher in Molecular Biology. 1969-1970 Dept. of Genetics, University of Washington (Seattle, USA), Research Associate . 1971-1987 Dept. of Genetics and Molecular Biology, University of Rome, “Sapienza”, Assistant Professor and Associate Professor in Molecular Biology. 1988- present Dept. of Genetics and Molecular Biology, University of Rome “Sapienza” Full Professor in Molecular Biology. 1992-2002 Director of the Centro Studi Acidi Nucleici (CNR, Rome). 2001-2006 Coordinator of the PhD Program in Pasteurian Sciences (University of Rome) . 2002-2010 Scientific Director of the Foundation “Institut Pasteur-Cenci Bolognetti”. Associate Editor of Physics of Life Reviews (Elsevier), Member of EMBO (European Molecular Biology Organization), of ISSOL (Intl. Soc for the Origin of Life), of AABMB (Am.Ass. Biochem. Mol. Biol.) The research activity has been consistently focused on the study of the structure and of the function of genetic materials, in both prokaryotes and eukaryotes. Contributions have been made in the fields of DNA topology and of topology-related functions, in the genetics, biophysics and biochemistry of gene expression. Special interest was devoted to kinetic and topological aspects of gene regulation. Another line of research dealt with the chemistry of DNA and has been centered on both the development of chemical sequencing technologies and on the study of the plausible prebiotic routes to the evolution of informational macromolecules. This latter approach has recently lead to the discovery of novel aspects of the mechanisms for the prebiotic synthesis of a whole new set of nucleic acids precursors. The study of novel chemical mechanisms in an origin-of-life scenario is at present one of the main interests of his research group. Overall, the results obtained in his research in biochemistry/molecular genetics have dealt with the topics: Telomeres, Chromatin, Nucleosomes, Gene regulation. DNA topology and DNA topoisomerases. DNA sequencing technologies and DNA conformation studies. RNA polymerases. |