Tel :+39 06 49913632
Fax :
nicole.balasco@cnr.it
Fax :
nicole.balasco@cnr.it
NICOLE BALASCO
Researcher
Institute of Molecular Biology and Pathology - National Research Council
Department of Chemistry, Sapienza University of Rome, P. le A. Moro 5, 00185 Rome
Building S. Cannizzaro (CU014), third floor, room 306
Researcher
Institute of Molecular Biology and Pathology - National Research Council
Department of Chemistry, Sapienza University of Rome, P. le A. Moro 5, 00185 Rome
Building S. Cannizzaro (CU014), third floor, room 306
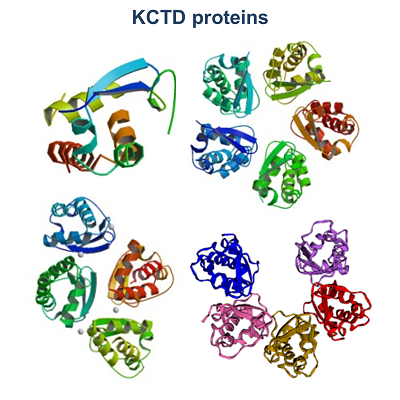
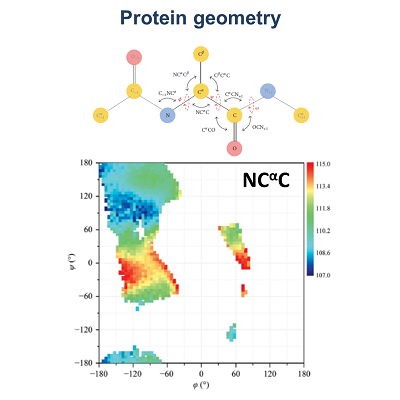
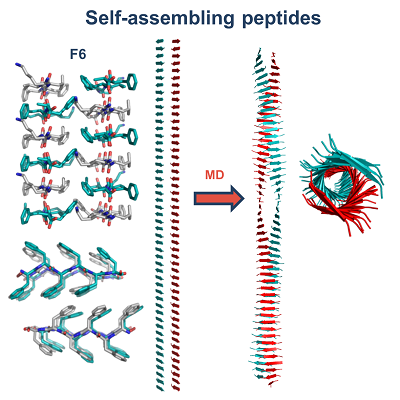
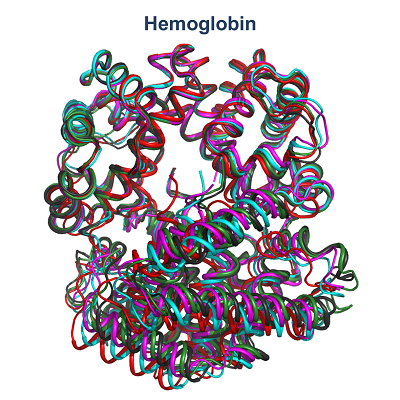
| My research activities are focused on the structural characterization of biomolecules endowed with a different level of molecular/structural complexity ranging from peptides to macromolecular complexes. These analyses are performed by combining computational and experimental approaches on a variety of diversified peptide/protein systems. In particular, my expertise includes the biophysical characterization of peptides/proteins in solution by means of spectroscopic techniques (circular dichroism, fluorescence) and light scattering, the structural characterization of proteins by X-ray crystallography, data mining in databases of peptide/protein structures, statistical analyses, and finally molecular design, molecular modeling, and molecular dynamics simulations. Principal research lines in which I am currently involved can be summarized as follow: • KCTD proteins • Protein geometry • Biomaterials based on self-assembling peptides in cross-beta structures • Structural bases of protein thermostability – manipulation of thermostable proteins • HCV proteins • Hemoglobin - structural transition of hemoglobins • Thrombin interaction with aptamers • Interaction of proteins with denaturants • Epitope recognition by MHC • Epidemiological analyses on COVID-19 disease • Mutational analyses on SARS-CoV-2 proteins |
| 1. Esposito L, Balasco N, Vitagliano L. Alphafold Predictions Provide Insights into the Structural Features of the Functional Oligomers of All Members of the KCTD Family. Int J Mol Sci. 2022 Nov 1;23(21):13346. doi: 10.3390/ijms232113346. 2. Balasco N, Esposito L, De Simone A, Vitagliano L. Local Backbone Geometry Plays a Critical Role in Determining Conformational Preferences of Amino Acid Residues in Proteins. Biomolecules. 2022 Aug 26;12(9):1184. doi:10.3390/biom12091184. 3. Balasco N, Damaggio G, Esposito L, Villani F, Berisio R, Colonna V, Vitagliano L. A global analysis of conservative and non-conservative mutations in SARS-CoV-2 detected in the first year of the COVID-19 world-wide diffusion. Sci Rep. 2021 Dec 30;11(1):24495. doi: 10.1038/s41598-021-04147-1. 4. Troisi R, Balasco N, Autiero I, Vitagliano L, Sica F. Exosite Binding in Thrombin: A Global Structural/Dynamic Overview of Complexes with Aptamers and Other Ligands. Int J Mol Sci. 2021 Oct 6;22(19):10803. doi:10.3390/ijms221910803. 5. Balasco N, Alba J, D'Abramo M, Vitagliano L. Quaternary StructureTransitions of Human Hemoglobin: An Atomic-Level View of the Functional Intermediate States. J Chem Inf Model. 2021 Aug 23;61(8):3988-3999. doi:10.1021/acs.jcim.1c00315. 6. Balasco N, Diaferia C, Morelli G, Vitagliano L, Accardo A. Amyloid-Like Aggregation in Diseases and Biomaterials: Osmosis of Structural Information. Front Bioeng Biotechnol. 2021 Mar 3;9:641372. doi: 10.3389/fbioe.2021.641372. 7. Balasco N, Esposito L, Vitagliano L. Factors affecting the amplitude of the t angle in proteins: a revisitation. Acta Crystallogr D Struct Biol. 2017 Jul 1;73(Pt 7):618-625. doi:10.1107/S2059798317007793. 8. Balasco N, Barone D, Sandomenico A, Ruggiero A, Doti N, Berisio R, Ruvo M, Vitagliano L. Structural Versatility of Hepatitis C Virus Proteins: Implications for the Design of Novel Anti-HCV Intervention Strategies. Curr Med Chem. 2017 Nov 24;24(36):4081-4101. doi: 10.2174/0929867324666170508105544. 9. Balasco N, Barone D, Vitagliano L. Structural conversion of the transformer protein RfaH: new insights derived from protein structure prediction and molecular dynamics simulations. J Biomol Struct Dyn. 2015;33(10):2173-9. doi:10.1080/07391102.2014.994188. 10. Balasco N, Esposito L, De Simone A, Vitagliano L. Role of loops connecting secondary structure elements in the stabilization of proteins isolated from thermophilic organisms. Protein Sci. 2013 Jul;22(7):1016-23. doi: 10.1002/pro.2279. |
| Since I recently moved from IBB CNR (Naples) to Rome, most of my research activities are based on collaborations with my previous group. |
| Dr. Luigi Vitagliano, Dr. Luciana Esposito, Dr. Alessia Ruggiero, Dr. Antonella Paladino, Dr. Ida Autiero - IBB CNR Naples Dr. Giovanni Smaldone - IRCCS SDN SYNLAB Naples Prof. Mario Guarracino - University of Cassino Dr. Amarinder Singh Thind - University of Wollongong, Australia Dr. Vincenza Colonna - IGB CNR Naples Dr. Cinzia Giannini - IC CNR Bari Prof. Alfonso De Simone - Imperial College Londra, University Federico II Naples Prof. Giuseppe Graziano - University of Sannio Benevento Prof. Filomena Sica, Dr. Romualdo Troisi - University Federico II Naples Prof. Giancarlo Morelli, Prof. Antonella Accardo, Dr. Carlo Diaferia - University Federico II Naples Prof. Marco D’Abramo - Sapienza University of Rome |
| EDUCATION 2015 December: Ph.D. in Molecular and Cellular Biotechnology, Second University of Naples, Thesis: “Analysis of the main determinants of protein three-dimensional structures”. 2011 October: Master degree in Chemistry (Curriculum: Physical-Chemistry) at Sapienza University of Rome. Thesis: “Oxidation of human albumin: a structural and spectroscopic study”. Final result: 110/110 cum laude. 2009 September: Bachelor of Chemistry at Sapienza University of Rome. Thesis : “Human serum albumin: correlation between stability and binding”. Final result: 110/110 cum laude. 2006 July: High school diploma – Liceo scientifico “Leonardo da Vinci”, Fiumicino (Rome). Final result: 100/100. PROFESSIONAL EXPERIENCE 2022 February - Present: Researcher at Institute of Molecular Biology and Pathology IBPM CNR, Rome. 2012 October - 2022 January: Fellowship at Institute of Biostructures and Bioimaging IBB CNR, Naples. 2015 April - May: Research activity at Division of Molecular Biosciences, Imperial College, London, UK. 2012 June - September: Intern (Regulatory and Medical Affairs) at Johnson & Johnson S.p.A., Rome. TECHNICAL COMPETENCIES Advanced skills in Structural and Computational Biology: - Biophysical characterization of peptides/proteins in solution by means of spectroscopic techniques (circular dichroism, fluorescence) and light scattering. - Structural characterization of proteins by X-ray crystallography. - Data mining in databases of protein/peptide structures and statistical analyses. - Molecular design, molecular modeling, molecular dynamics simulations. PUBLICATIONS H-index = 19 (SCOPUS) 64 published papers (33 as first/co-first author, 13 as corresponding author) 6 preprints RESEARCH PROJECT PI Principal investigator of the CNR unit in the research project by the Italian Ministry for Universities and Research (MUR) PRIN (Progetto di Ricerca di Rilevante Interesse Nazionale), Title: Biomaterials from peptide self-assembling generated by mimicking protein amyloid-like structures – Code: 2022TSLMHR, Validity: 16/10/2023 to 16/10/2025. PI in the following CINECA ISCRA projects: IsCa3 AF-Koli (2022-2023), IsC95 AntHbs (2021-2022), IsC82 ArgSens (2020-2021), IsC70 HsHbDYN (2019-2020), IsC52 KCTDOLI (2017-2018). SCIENTIFIC CONFERENCE ORGANIZATION 2025 september 2-5: Member of the Scientific Committee of the "51st Meeting of the Italian Crystallographic Association (AIC)", Firenze. 2019 August 29 -September 3: Member of the Organizing Committee of “AIC International Crystallographic School 2019 CIF”, Naples. 2019 September 4-7: Member of the Organizing Committee of “Fifth Meeting of the Italian and Spanish Crystallographic Associations MISCA V”, Naples. TEACHING ACTIVITY 2024-2025: Teacher of Organic Chemistry at Scuola di Alta Formazione e Studio of ICPAL (Istituto centrale per la patologia degli archivi e del libro) - Ministero della Cultura, Roma. 2014 - 2019: Teaching Assistant of Chemistry (Engineering course) at University of Sannio, Benevento. 2012 - 2013 : Teacher of Chemistry at Alpha Test S.r.l. OTHER ACTIVITIES Co-author of 15 crystallographic protein structures deposited in the Protein Data Bank. Co-author of the website (https://alphafold.ibb.cnr.it/) that reports a database of AlphaFold predicted structures of KCTD oligomers. Co-author of the website RECOVER-COVID-19 (http://health.ibb.cnr.it/recover) that reports epidemiological and mutational analyses on COVID-19 disease and SARS-CoV-2 proteins, respectively. Co-author of the website QUIPROQUA for the analysis of the protein fine structure in quality assessment. |