Fax :+390649912343
patrizia.somma@cnr.it
Researcher
Sapienza University of Rome
Institute of Molecular Biology and Pathology CNR
Dipartimento di Biologia e Biotecnologie ex Edificio di Genetica I Piano stanza 2-25 P.le A.Moro 5 00185 Roma
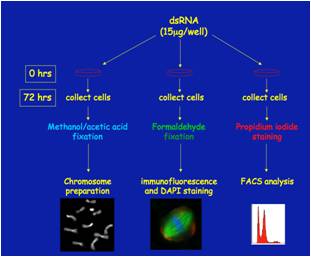
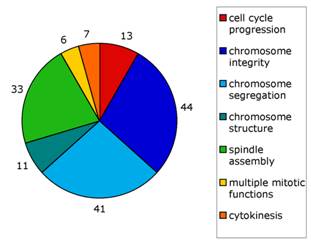
We use Drosophila melanogaster as a model organism to investigate complex processes such as cell division. The sophisticated genetics and the ease of RNA interference (RNAi) in Drosophila offer unique possibilities to study the process of mitosis. As most Drosophila mitotic genes are highly conserved, our results will help to understand the molecular mechanisms of cell division in vertebrates, including humans. Indeed, these approaches have enabled us to construct a gene expression signature (the DM signature, patent pending) using the human orthologues of 108 Drosophila mitotic genes. These 108 genes constitute an excellent signature to predict breast cancer outcome, as their increased expression negatively correlates with patient survival (Somma et al, 2010; Damasco et al., 2011)
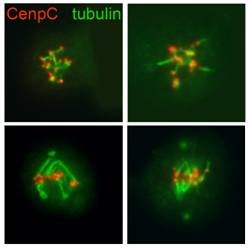
Our goup has recently undertaken an RNAi-based screen that has led to the identification of several proteins required for chromosome-driven microtubule (MT)-formation and/or involved in regulation of MT dynamics at kinetchores. We are currently investigating the role of these factors in MT spindle assembly and function.
We use different experimental approaches to gain insight into the functions of mitotic proteins: cytological analysis using fluorescence microscopy, in vivo analysis of cell division, biochemical techniques to identify complexes containing the mitotic proteins of interest. |
1)Damasco C., Lembo A., Somma M.P., Gatti M., Di Cunto F., Provero P. A signature inferred from Drosophila mitotic genes predicts survival of breast cancer patients. PLoS One. 2011 Feb 28;6(2):e14737.
2)Bucciarelli E., Pellacani C., Naim V., Palena A., Gatti M., Somma M.P. The Drosophila Dgt6 protein interacts with g-tubulin, Msps/XMAP215 and Ndc80 to promote kinetochore-driven MT formation. Curr. Biol. 2009 Nov 17;19(21):1839-45.
3)Somma M.P., Ceprani F., Bucciarelli E., Naim V., De Arcangelis V., Piergentili R., Palena A., Ciapponi L., Giansanti M.G., Pellacani C., Petrucci R., Cenci G., Vernì F., Fasulo B., Goldberg M.L., Di Cunto F., Gatti M. Identification of Drosophila mitotic genes by combining co-expression analysis and RNA interference. PLoS Genet. Jul 18;4(7):e1000126, (2008).
4)Yu J., Fleming S.L., Williams B., Williams E.V., Li Z., Somma P., Rieder C.L., Goldberg M.L. Greatwall kinase: a nuclear protein required for proper chromosome condensation and mitotic progression in Drosophila. J Cell Biol. 164(4):487-92, (2004).
5)Vernì F., Somma M.P., Gunsalus K.C., Bonaccorsi S., Belloni G., Goldberg M.L., Gatti M. Feo, the Drosophila homolog of PRC1, is required for central-spindle formation and cytokinesis. Curr Biol. 14(17):1569-75, (2004).
6)Somma M.P., Fasulo B., Siriaco G., Cenci G. Chromosome condensation defects in barren RNA-interfered Drosophila cells. Genetics 165(3):1607-11, (2003).
7)Somma M.P., Fasulo B., Cenci G., Cundari E., Gatti M. Molecular dissection of cytokinesis by RNA interference in Drosophila cultured cells. Mol Biol Cell. 13(7):2448-60, (2002).
8)Stapleton G., Somma M.P., Lavia P. Cell type-specific interactions of transcription factors with a housekeeping promoter in vivo. Nucleic Acids Res., 21: 2465-2471, (1993).
9) Somma M.P., Gambino I., Lavia P. Transcription factors binding to the mouse HTF9 housekeeping promoter differ between cell types. Nucleic Ac. Res.19: 4451-4458, (1991).
10)Somma M.P., Pisano C., Lavia P. The housekeeping promoter from the mouse CpG island HTF9 contains multiple protein-binding elements that are functionally redundant. Nucleic Ac. Res., 19: 2817-2824, (1991). |
- Elisabetta Bucciarelli, CNR post-doctoral research fellow - Claudia Pellacani, post-doctoral research fellow - Fioranna Renda, PhD student - Antonella Palena, CNR Staff |
Within IBPM Maurizio Gatti, Dip. Biologia e Biotechnologie “Charles Darwin”, Sapienza
With other institutions Michael L. Goldberg, Dept. of Molecular Biology & Genetics, Cornell University Ithaca, NY, USA |
Place and date of birth: Rome 23 June 1960
Education: - 1991: PhD in Evolutionary Biology, Sapienza University, Rome - 1986: Laurea in Biology cum Laude, Sapienza University, Rome
Research Positions: - 2001 – present: CNR Research Scientist (Permanent appointment) at IBPM, Rome - 1996-2001: CNR Research Scientist (5-year position ex-art.36 CNR) at CNR Center of Evolutionary Genetics, Rome - 1992- 1996: Post-doctoral research fellow at CNR Center of Evolutionary Genetics, Rome - April-May 1991: Visiting fellow, Academic research contract, AFRC-Centre for Genome Research, University of Edinburgh (UK) (Director Prof. R. Lathe) |