Fax :+39064440062
veronica.morea@cnr.it
Senior Researcher
Istitute of Molecular Biology and Pathology (IBPM)
National Research Council (CNR)
Department of Biochemical Sciences -A. Rossi Fanelli. - Sapienza University of Rome P.le A. Moro 5 - 00185 Roma Edificio di Chimica Biologica Piano -1, stanza S26
I) Structural analysis of protein and tRNA molecules and investigation of relationships between sequences, structures and functions; II) Design of proteins and peptides with desired properties III) Prediction of protein structure and function IV) Biodiversity and evolutionary studies at the molecular level
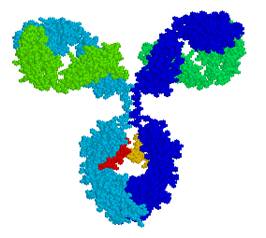
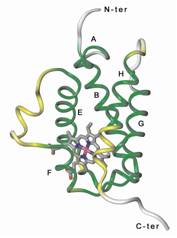
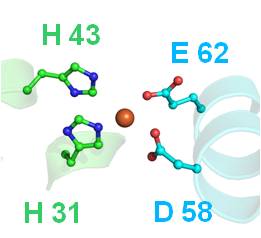
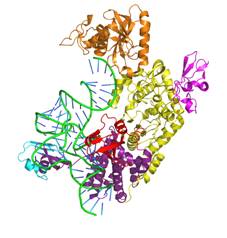
Main projects: 1) Design of nanoparticles based on proteins belonging tot he ferritin family, aimed at selectively bind to and be internalized by target cells and tissues, for diagnostic and therapeutic (“theragnostic”) applications; 2) Design of humanized variants of non-human antibodies for clinical appplications; 3) Structural analysis of proteins involved in angiogenesis (e.g., VEGFR-1, alpha5beta1 integrin, neuropilin.1, etc.) and design of pro- or anti-angiogenic peptides; 4) Structural analysis of tRNAs and of proteins interacting with them aimed at understanding the molecular basis of the pathological effects caused by point mutations, and designing peptides able to antagonize these defects. |
1.Vannucci L, Falvo E, Fornara M, Di Micco P, Benada O, Krizan J, Svoboda J, Hulikova-Capkova K, Morea V, Boffi A, Ceci P. “Selective targeting of melanoma by PEG-masked protein-based multifunctional nanoparticles” Int J Nanomed (2012) 7: 1489 – 1509
2.Perli E, Giordano C, Tuppen HA, Montopoli M, Montanari A, Orlandi M, Pisano A, Catanzaro D, Caparrotta L, Musumeci B, Autore C, Morea V, Di Micco P, Campese AF, Leopizzi M, Gallo P, Francisci S, Frontali L, Taylor RW, d'Amati G. Isoleucyl-tRNA synthetase levels modulate the penetrance of a homoplasmic m.4277T>C mitochondrial tRNA(Ile) mutation causing hypertrophic cardiomyopathy. Hum Mol Genet (2012) 21: 85-100.
3.Soro S, Orecchia A, Morbidelli L, Lacal PM, Morea V, Ballmer-Hofer K, Ruffini F, Ziche M, D'Atri S, Zambruno G, Tramontano A, Failla CM. “A proangiogenic peptide derived from vascular endothelial growth factor receptor-1 acts through alpha5beta1 integrin.” (2008) Blood, 111: 3479-3488
4.Vogel C, Morea V. “Duplication, divergence and formation of novel protein topologies” (2006) Bioessays, 28: 973-978. Review
5.Tramontano A, Morea V. “Assessment of homology-based predictions in CASP5” (2003) Proteins – Structure, Function and Genetics, 53 Suppl. 6: 352-368.
6.Donini M*, Morea V*, Desiderio A, Pashkoulov D, Villani ME, Tramontano A, Benvenuto E. “Engineering Stable Cytoplasmic Intrabodies with Designed Specificity” (2003) J. Mol. Biol., 330: 323-332.
7.Tramontano A, Leplae R, Morea V. “Analysis and assessment of comparative modeling predictions in CASP4” (2001) Proteins – Structure, Function and Genetics, Suppl. 5: 22-38.
8.Morea V, Lesk AM, Tramontano A. “Antibody modeling: implications for engineering and design.” (2000) Methods – A companion to Methods in Enzymology, 20: 267-279. Review
9.Morea V, Tramontano A, Rustici M, Chothia C, Lesk AM. “Conformations of the third hypervariable region in the VH domain of immunoglobulins” (1998) J. Mol. Biol., 275: 269-294.
10.Morea V, Tramontano A, Rustici M, Chothia C, Lesk AM. “Antibody structure, prediction and redesign” (1997) Biophys. Chem., 68(1-3): 9-16. |
Patrizio Di Micco, PhD student in Biochemistry at the University of Rome “Sapienza” Ferdinando Spagnolo, student of the Master in Bioinformatics of the University of Rome “Sapienza” |
CNR, Institute of Molecular Biology and Pathology (IBPM): Pierpaolo Ceci Gianni Colotti Adele Di Matteo Andrea Ilari Carmelinda Savino
University of Rome “Sapienza”, Department of Biochemical Sciences “A. Rossi Fanelli”: Andrea Bellelli Alberto Boffi Maurizio Brunori Emilia Chiancone Roberta Chiaraluce Valerio Consalvi Beatrice Vallone
University of Rome “Sapienza”, Department of Biology and Biotechnology “Charles Darwin”: Silvia Francisci Laura Frontali
University of Rome “Sapienza”, Department of Experimental Medicine: Giulia D'Amati Carla Giordano
Istituto Dermopatico dell'Immacolata (IDI)-IRCCS, Rome: Stefania D'Atri Cristina M. Failla Pedro M. Lacal Giovanna Zambruno
Istituto Nazionale Tumori Regina Elena (IRE)-IRCCS, Rome: Rita Falcioni Michele Milella
Institute of Animal Physiology and Genetics of the Academy of Sciences of the Czech Republic, Prague, Czech Republic: Luca Vannucci
University of L'Aquila, Department of Pure and Applied Biology: Francesco Angelucci Rodolfo Ippoliti
University of Verona, Department of Pathology and Diagnostics Marco Colombatti Fracasso Giulio
CNR, Institute of Biology and Biotechnology (IBBA): Aldo Ceriotti
Italian National Agency for New Technologies, Energy and Sustainable Economic Development (ENEA), Casaccia Research Centre, Cesano (Rome): Eugenio Benvenuto Marcello Donini Alessandra Pasquo Maria Elena Villani
St. George's University of London, UK Julian Ma
University of Naples “Parthenope”: Romina Oliva |
Working experience
From 2001: Research Scientist at the Institute of Molecular Biology and Pathology (IBPM) of the National Research Council (CNR). 2000-2001: Research Scientist at the Medical Research Council (MRC), Laboratory of Molecular Biology (LMB), Cambridge (UK) in Dr. Cyrus Chothia’s Bioinformatics group 1998-2000: Marie-Curie Research Fellow (Biotechnology Programme) at the MRC-LMB, Cambridge (UK) in Dr. Cyrus Chothia’s Bioinformatics group 1996-1998: Research Scientist at the University “G. D’Annunzio”, Chieti, Italy
Education
10/07/1997: Ph. D. at the Institute of Research in Molecular Biology (IRBM) “P. Angeletti”, Pomezia (Rome), Italy, in Prof. Anna Tramontano’s Bioinformatics group. Thesis: “Analysis and prediction of protein structures and of protein-ligand interactions with computational and kinetic methods: implications for the design of therapeutic agents.” 20/07/1993: Degree in Pharmacy cum laude at the University of Rome “Sapienza” (IT). Experimental thesis: “Peptidyl-methyl-ketones as reversibile substrate-analogue inhibitors of cysteine proteases.” 5/11/1992: Degree in Medicinal Chemistry cum laude at the University “Sapienza”, Rome, Italy. Experimental thesis: “Selective inhibitors of cysteine proteases: synthesis and kinetic characterization.”
Awards
1993: The thesis for the degree in Medicinal Chemistry has been awarded by the BRACCO pharmaceutical company.
Teaching
2012: “Structure-based design of proteins and peptides endowed with desired properties”, University of Rome “Sapienza”, Master in Bioinformatics 2011: “Bioinformatics Applications in Structural and Molecular Biology”, University of L’Aquila, Ph. D. course 2010: “Bioinformatics Applications in Molecular and Structural biology”, University of Rome “Sapienza”, advanced course for CNR personnel 2005: “Bioinformatics Applications in Molecular and Structural biology”, University of Rome “Sapienza”, advanced course for CNR personnel 2005: “Bioinformatics for Molecular and Cellular Biologists”, University of Rome “Sapienza”, Ph. D. course 2004: “Bioinformatics”, University of Rome “Sapienza”, advanced course for CNR personnel 10/1996-09/1997: “Analysis of pharmaceutical products” and “Preparation of pharmaceutical products with extractive and synthetic methods”, University “G. D’Annunzio” of Chieti (IT), degree in Medicinal Chemistry. |